All in One View
Content from Before we start
Last updated on 2025-02-26 | Edit this page
Overview
Questions
- What is Python and why should I learn it?
Objectives
- Present motivations for using Python.
- Organize files and directories for a set of analyses as a Python project.
- How to work with Jupyter Notebook.
- Know where to find help.
This overview have been adapted from the Data Carpentry Python Lessons with Ecological Data.
Why Python?
Python is rapidly becoming the standard language for data analysis,
visualization and automated workflow building. It is a free and
open-source software that is relatively easy to pick up by new
programmers and is available on multiple operating systems. In addition,
with Python packages such as Jupyter one can keep an
interactive code journal of analysis - this is what we’ll be using in
the workshop. Using Jupyter notebooks allows you to keep a record of all
the steps in your analysis, enabling reproducibility and ease of code
sharing.
Another advantage of Python is that it is maintained by a large user-base. Anyone can easily make their own Python packages for others to use. Therefore, there exists a very large codebase for you to take advantage of for your neuroimaging analysis; from basic statistical analysis, to brain visualization tools, to advanced machine learning and multivariate methods!
Knowing your way around Anaconda and Jupyter
The Anaconda distribution of Python includes a lot of its popular packages, such as the IPython console, and Jupyter Notebook. Have a quick look around the Anaconda Navigator. You can launch programs from the Navigator or use the command line.
The Jupyter notebook is an open-source web application that allows you to create and share documents that allow one to create documents that combine code, graphs, and narrative text. If you have not worked with Jupyter notebooks before, you might start with the tutorial from Jupyter.org called “Try Classic Notebook”.
Anaconda also comes with a package manager called conda, which makes it easy to install and update additional packages.
Organizing your working directory
Using a consistent folder structure across your projects will help you keep things organized, and will also make it easy to find/file things in the future. This can be especially helpful when you have multiple projects. In general, you may wish to create separate directories for your scripts, data, and documents.
This lesson uses the following folders to keep things organized:
data/: Use this folder to store your raw data. For the sake of transparency and provenance, you should always keep a copy of your raw data. If you need to cleanup data, do it programmatically (i.e. with scripts) and make sure to separate cleaned up data from the raw data. For example, you can store raw data in files./data/raw/and clean data in./data/clean/.code/: Use this folder to store your (Python) scripts for data cleaning, analysis, and plotting that you use in this particular project. This folder contains all the Jupyter notebooks used in the lesson.files/: Use this folder to store any miscellaneous outlines, drafts, and other text. This folder contains a set of Powerpoint slides that will be used for parts of the lesson.
How to learn more after the workshop?
The material we cover during this workshop will give you an initial taste of how you can use Python to analyze MRI data for your own research. However, you will need to learn more to do advanced operations such as working with specific MRI modalities, cleaning your dataset, using statistical methods, or creating beautiful graphics. The best way to become proficient and efficient at Python, as with any other tool, is to use it to address your actual research questions. As a beginner, it can feel daunting to have to write a script from scratch, and given that many people make their code available online, modifying existing code to suit your purpose might make it easier for you to get started.
Seeking help
- check under the Help menu
- type
help() - type
?objectorhelp(object)to get information about an object - Python documentation
- NiBabel documentation
Finally, a generic Google or internet search “Python task” will often either send you to the appropriate module documentation or a helpful forum where someone else has already asked your question.
I am stuck… I get an error message that I don’t understand. Start by googling the error message. However, this doesn’t always work very well, because often, package developers rely on the error catching provided by Python. You end up with general error messages that might not be very helpful to diagnose a problem (e.g. “subscript out of bounds”). If the message is very generic, you might also include the name of the function or package you’re using in your query.
You can also check Stack Overflow. Most questions have already been answered, but the challenge is to use the right words in the search to find the answers.
Another helpful resource specific to the neuroimaging community is NeuroStars.
- Python is an open source and platform independent programming language.
- Jupyter Notebook is a great tool to code in and interact with Python.
- With the large Python community it is easy to find help on the internet.
Content from From the scanner to our computer
Last updated on 2025-02-26 | Edit this page
Overview
Questions
- What are the main MRI modalities?
- What’s the first step necessary to start working with MRI data?
Objectives
- Understand how different MRI modalities differ and what each one represents
- Become familiar with converting MRI data from DICOM to NIfTI
Types of MR scans
Anatomical
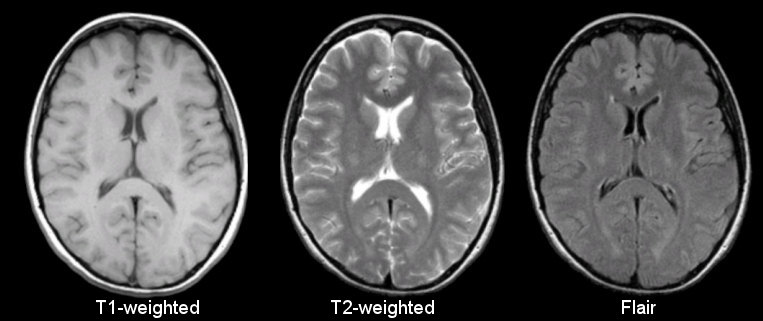
Sourced from https://case.edu/med/neurology/NR/MRI%20Basics.htm
- 3D image of anatomy (i.e., shape, volume, cortical thickness, brain region)
- can differentiate tissue types
Functional
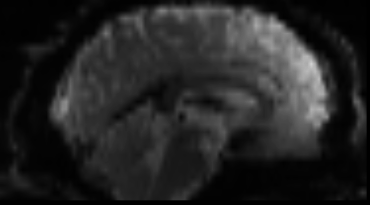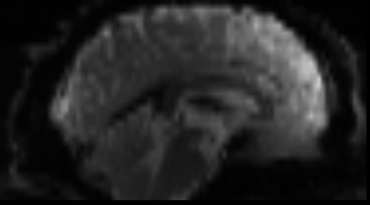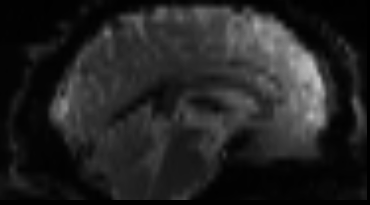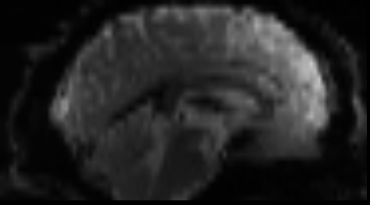
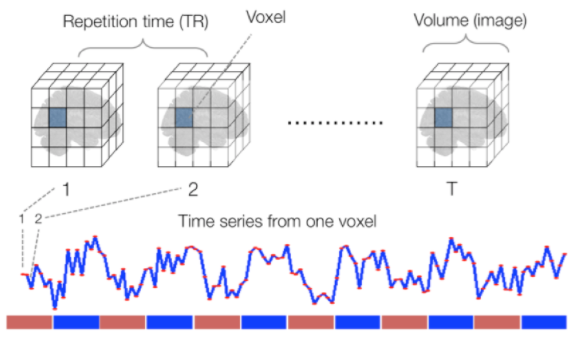
Sourced from Wagner and Lindquist, 2015
- tracks the blood oxygen level-dependant (BOLD) signal as an analogue of brain activity
- 4D image (x, y, z + time)
Diffusion

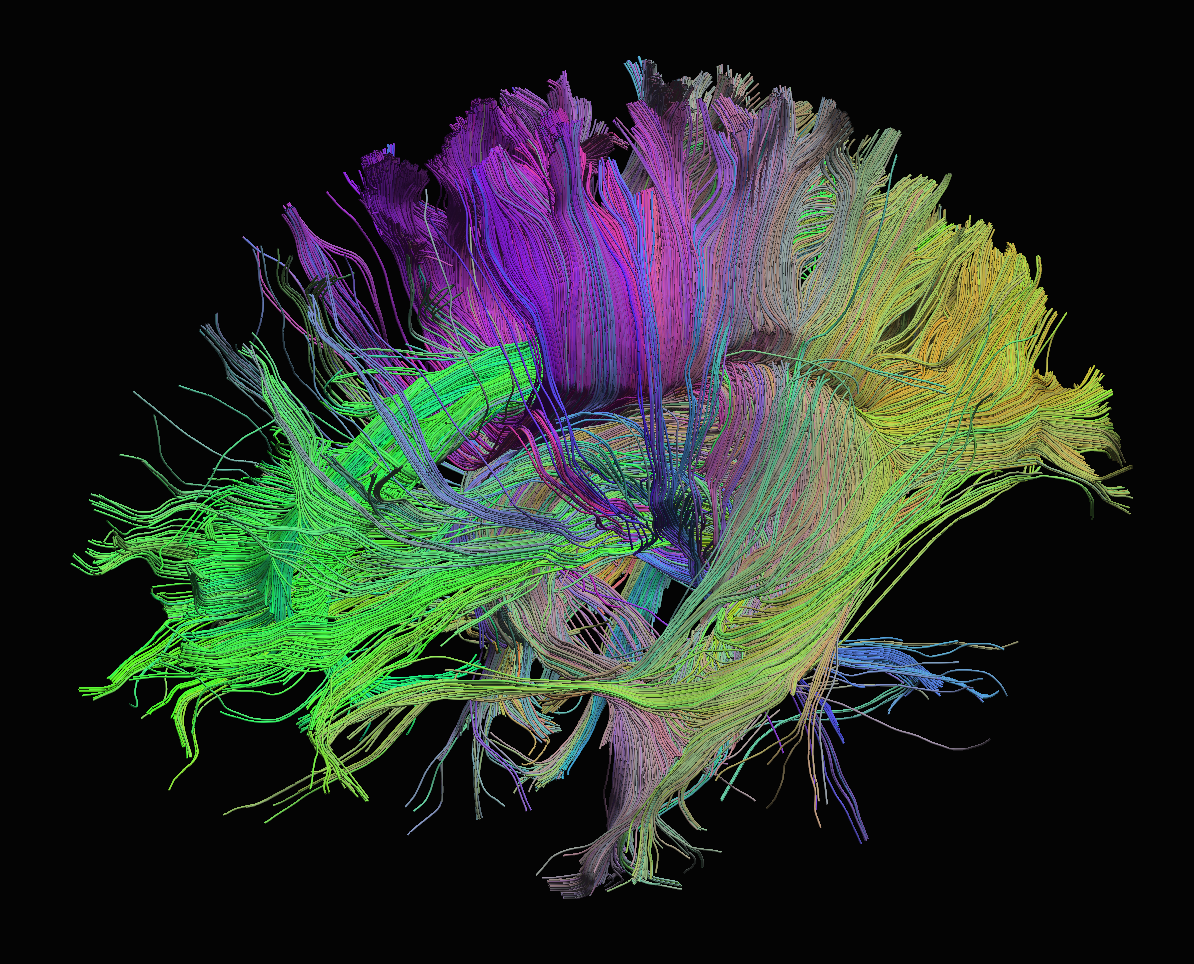
Sourced from http://brainsuite.org/processing/diffusion/tractography/
- measures diffusion of water in order to model tissue microstructure
- 4D image (x, y, z + direction of diffusion)
- need parameters about the strength of the diffusion “gradient” and
its direction (
.bvaland.bvec)
Neuroimaging file formats
Common file formats:
| Format Name | File Extension | Origin/Group |
|---|---|---|
| DICOM | none (*) | ACR/NEMA Consortium |
| Analyze |
.img/.hdr
|
Analyze Software, Mayo Clinic |
| NIfTI |
.nii (**) |
Neuroimaging Informatics Technology Initiative |
| MINC | .mnc |
Montreal Neurological Institute |
| NRRD | .nrrd |
Gordon L. Kindlmann |
| MGH |
.mgz or .mgh
|
Massachusetts General Hospital |
(*) DICOM files typically have a .dcm file extension.
(**) Some files are typically compressed and their extension may
incorporate the corresponding compression tool extension
(e.g. .nii.gz).
From the MRI scanner, images are initially collected in the DICOM format and can be converted to these other formats to make working with the data easier.
Let’s download some example DICOM data to see what it looks like. This data was generously shared publicly by the Princeton Handbook for Reproducible Neuroimaging.
BASH
wget https://zenodo.org/record/3677090/files/0219191_mystudy-0219-1114.tar.gz -O ../data/0219191_mystudy-0219-1114.tar.gz
mkdir -p ../data/dicom_examples
tar -xvzf ../data/0219191_mystudy-0219-1114.tar.gz -C ../data/dicom_examples
gzip -d ../data/dicom_examples/0219191_mystudy-0219-1114/dcm/*dcm.gz
rm ../data/0219191_mystudy-0219-1114.tar.gz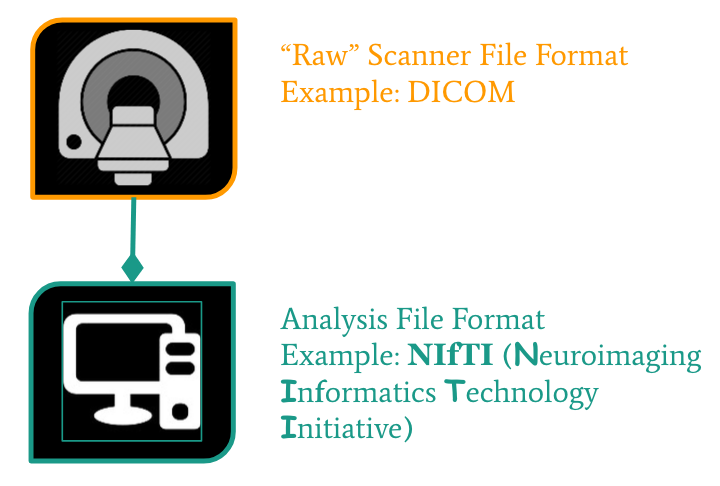
NIfTI is one of the most ubiquitous file formats for storing neuroimaging data. If you’re interested in learning more about NIfTI images, we highly recommend this blog post about the NIfTI format. We can convert our DICOM data to NIfTI using dcm2niix.
We can learn how to run dcm2niix by taking a look at its
help menu.
Converting DICOM to NIfTI
Convert the Princeton DICOM data to NIfTI
- MRI can capture anatomical (structural), functional, or diffusion features.
- A number of file formats exist to store neuroimaging data.
Content from Anatomy of a NIfTI
Last updated on 2025-02-26 | Edit this page
Overview
Questions
- How are MRI data represented digitally?
Objectives
- Introduce Python data types
- Load an MRI scan into Python and understand how the data is stored
- Understand what a voxel is
- Introduce MRI coordinate systems
- View and manipulate image data
In the last lesson, we introduced the NIfTI. We’ll cover a few details to get started working with them.
Reading NIfTI images
NiBabel is a Python package for reading and writing neuroimaging data. To learn more about how NiBabel handles NIfTIs, check out the Working with NIfTI images page of the NiBabel documentation, from which this episode is heavily based.
First, use the load() function to create a NiBabel image
object from a NIfTI file. We’ll load in an example T1w image from the
zip file we just downloaded.
PYTHON
t1_img = nib.load("../data/dicom_examples/nii/0219191_mystudy-0219-1114_anat_ses-01_T1w_20190219111436_5.nii.gz")Loading in a NIfTI file with NiBabel gives us a special
type of data object which encodes all the information in the file.Each
bit of information is called an attribute in Python’s
terminology. To see all these attributes, type t1_img.
followed by Tab. There are three main attributes that we’ll
discuss today:
1. Header: contains metadata about the image, such as image dimensions, data type, etc.
OUTPUT
<class 'nibabel.nifti1.Nifti1Header'> object, endian='<'
sizeof_hdr : 348
data_type : b''
db_name : b''
extents : 0
session_error : 0
regular : b''
dim_info : 0
dim : [ 3 57 67 56 1 1 1 1]
intent_p1 : 0.0
intent_p2 : 0.0
intent_p3 : 0.0
intent_code : none
datatype : uint8
bitpix : 8
slice_start : 0
pixdim : [1. 2.75 2.75 2.75 1. 1. 1. 1. ]
vox_offset : 0.0
scl_slope : nan
scl_inter : nan
slice_end : 0
slice_code : unknown
xyzt_units : 2
cal_max : 0.0
cal_min : 0.0
slice_duration : 0.0
toffset : 0.0
glmax : 0
glmin : 0
descrip : b''
aux_file : b''
qform_code : mni
sform_code : mni
quatern_b : 0.0
quatern_c : 0.0
quatern_d : 0.0
qoffset_x : -78.0
qoffset_y : -91.0
qoffset_z : -91.0
srow_x : [ 2.75 0. 0. -78. ]
srow_y : [ 0. 2.75 0. -91. ]
srow_z : [ 0. 0. 2.75 -91. ]
intent_name : b''
magic : b'n+1't1_hdr is a Python dictionary.
Dictionaries are containers that hold pairs of objects - keys and
values. Let’s take a look at all the keys. Similar to
t1_img in which attributes can be accessed by typing
t1_img. followed by Tab, you can do the same
with t1_hdr. In particular, we’ll be using a
method belonging to t1_hdr that will allow
you to view the keys associated with it.
OUTPUT
['sizeof_hdr',
'data_type',
'db_name',
'extents',
'session_error',
'regular',
'dim_info',
'dim',
'intent_p1',
'intent_p2',
'intent_p3',
'intent_code',
'datatype',
'bitpix',
'slice_start',
'pixdim',
'vox_offset',
'scl_slope',
'scl_inter',
'slice_end',
'slice_code',
'xyzt_units',
'cal_max',
'cal_min',
'slice_duration',
'toffset',
'glmax',
'glmin',
'descrip',
'aux_file',
'qform_code',
'sform_code',
'quatern_b',
'quatern_c',
'quatern_d',
'qoffset_x',
'qoffset_y',
'qoffset_z',
'srow_x',
'srow_y',
'srow_z',
'intent_name',
'magic']Notice that methods require you to include () at the end of them whereas attributes do not. The key difference between a method and an attribute is:
- Attributes are stored values kept within an object
- Methods are processes that we can run using the object. Usually a method takes attributes, performs an operation on them, then returns it for you to use.
When you type in t1_img. followed by Tab,
you’ll see that attributes are highlighted in orange and methods
highlighted in blue.
The output above is a list of keys you can use from
t1_hdr to access values. We can access the
value stored by a given key by typing:
Extract values from the NIfTI header
Extract the value of pixdim from t1_hdr
2. Data
As you’ve seen above, the header contains useful information that
gives us information about the properties (metadata) associated with the
MR data we’ve loaded in. Now we’ll move in to loading the actual
image data itself. We can achieve this by using the method
called t1_img.get_fdata().
OUTPUT
array([[[ 8.79311806, 8.79311806, 8.79311806, ..., 7.73574093,
7.73574093, 7.38328189],
[ 8.79311806, 9.1455771 , 8.79311806, ..., 8.08819997,
8.08819997, 8.08819997],
[ 9.1455771 , 8.79311806, 8.79311806, ..., 8.44065902,
8.44065902, 8.44065902],
...,
[ 8.08819997, 8.44065902, 8.08819997, ..., 7.38328189,
7.38328189, 7.38328189],
[ 8.08819997, 8.08819997, 8.08819997, ..., 7.73574093,
7.38328189, 7.38328189],
[ 8.08819997, 8.08819997, 8.08819997, ..., 7.38328189,
7.38328189, 7.03082284]],
[[ 8.79311806, 9.1455771 , 8.79311806, ..., 7.73574093,
7.38328189, 7.38328189],
[ 8.79311806, 9.1455771 , 9.1455771 , ..., 8.08819997,
7.73574093, 8.08819997],
[ 8.79311806, 9.49803615, 9.1455771 , ..., 8.44065902,
8.44065902, 8.44065902],
...,
[ 8.08819997, 8.08819997, 8.08819997, ..., 7.38328189,
7.38328189, 7.03082284],
[ 8.08819997, 8.08819997, 8.08819997, ..., 7.38328189,
7.38328189, 7.38328189],
[ 8.08819997, 8.08819997, 8.08819997, ..., 7.38328189,
7.38328189, 7.73574093]],
[[ 9.1455771 , 9.1455771 , 8.79311806, ..., 7.73574093,
7.38328189, 7.03082284],
[ 9.1455771 , 9.49803615, 9.1455771 , ..., 8.08819997,
7.73574093, 7.38328189],
[ 9.1455771 , 9.49803615, 9.1455771 , ..., 8.08819997,
8.08819997, 8.08819997],
...,
[ 8.08819997, 8.44065902, 8.44065902, ..., 7.73574093,
7.38328189, 7.38328189],
[ 8.44065902, 8.08819997, 8.44065902, ..., 7.38328189,
7.38328189, 7.38328189],
[ 8.08819997, 8.08819997, 8.08819997, ..., 7.38328189,
7.38328189, 7.38328189]],
...,
[[ 9.49803615, 9.85049519, 9.85049519, ..., 7.38328189,
7.38328189, 7.03082284],
[ 9.85049519, 9.85049519, 9.85049519, ..., 7.38328189,
7.38328189, 7.73574093],
[ 9.85049519, 9.85049519, 10.20295423, ..., 8.08819997,
8.08819997, 8.08819997],
...,
[ 9.49803615, 9.1455771 , 9.49803615, ..., 7.73574093,
7.73574093, 7.73574093],
[ 9.49803615, 9.49803615, 9.1455771 , ..., 7.73574093,
8.08819997, 7.73574093],
[ 9.49803615, 9.1455771 , 8.79311806, ..., 8.08819997,
7.73574093, 7.73574093]],
[[ 9.49803615, 9.85049519, 10.20295423, ..., 7.38328189,
7.38328189, 7.03082284],
[ 9.1455771 , 9.85049519, 9.1455771 , ..., 7.73574093,
7.38328189, 7.38328189],
[ 9.85049519, 9.85049519, 9.49803615, ..., 8.08819997,
7.73574093, 8.08819997],
...,
[ 9.49803615, 8.79311806, 9.1455771 , ..., 8.08819997,
7.38328189, 7.73574093],
[ 9.1455771 , 9.1455771 , 9.1455771 , ..., 7.73574093,
8.08819997, 7.73574093],
[ 8.79311806, 9.1455771 , 9.1455771 , ..., 7.73574093,
7.73574093, 7.38328189]],
[[ 9.1455771 , 8.79311806, 8.79311806, ..., 7.03082284,
7.03082284, 6.6783638 ],
[ 9.49803615, 9.85049519, 9.49803615, ..., 7.73574093,
7.73574093, 7.73574093],
[ 9.49803615, 9.49803615, 9.85049519, ..., 7.73574093,
8.08819997, 7.73574093],
...,
[ 8.79311806, 9.1455771 , 9.1455771 , ..., 8.08819997,
8.08819997, 8.08819997],
[ 9.1455771 , 9.1455771 , 9.1455771 , ..., 8.08819997,
8.08819997, 7.73574093],
[ 8.79311806, 9.1455771 , 9.1455771 , ..., 8.08819997,
7.73574093, 7.73574093]]])What type of data is this exactly? We can determine this by calling
the type() function on t1_data.
OUTPUT
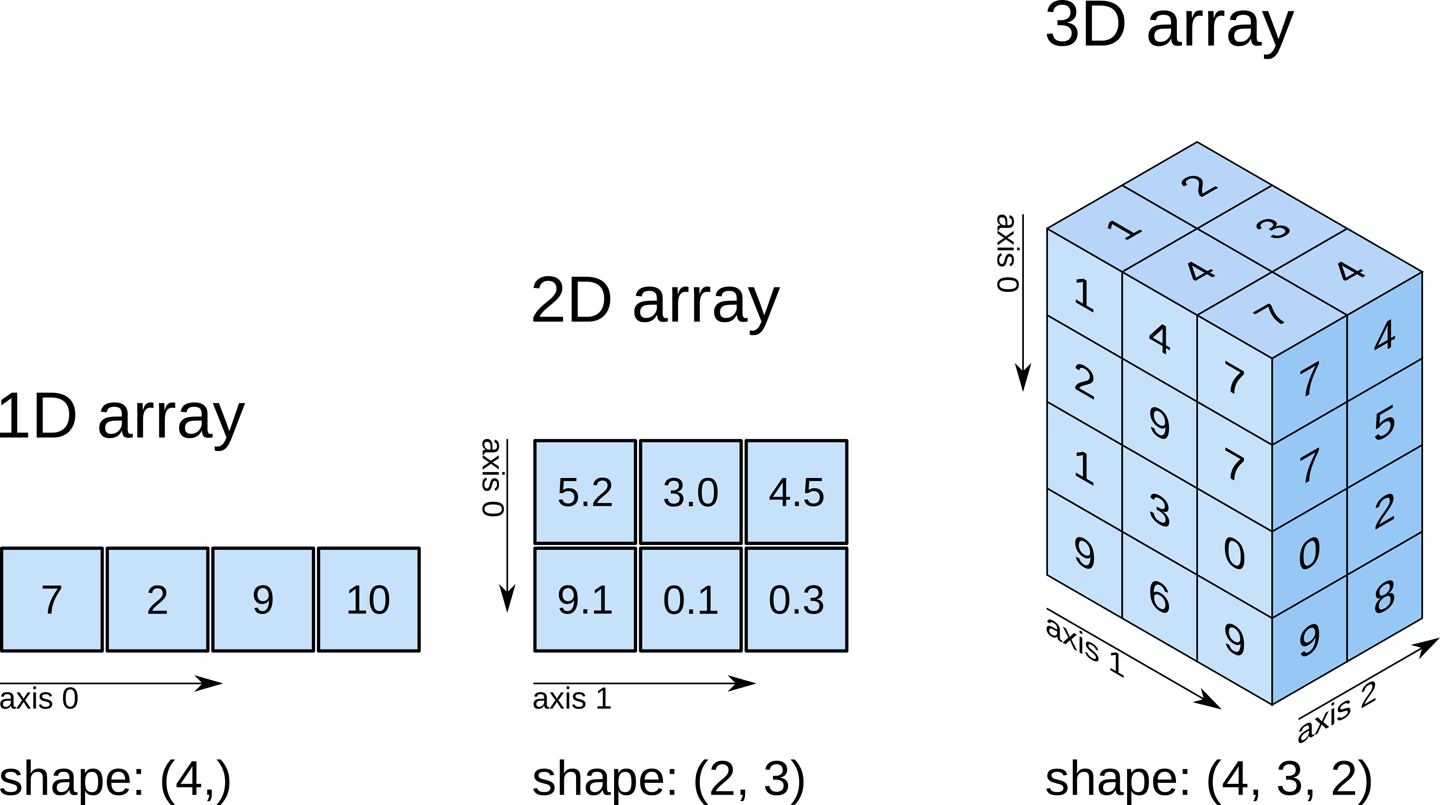
numpy.ndarrayThe data is a multidimensional array representing the image data. In Python, an array is used to store lists of numerical data into something like a table.
Check out attributes of the array
How can we see the number of dimensions in the t1_data
array? Once again, all the attributes of the array can be seen by typing
t1_data. followed by Tab.
Check out attributes of the array (continued)
While typical 2D pictures are made out of squares called
pixels, a 3D MR image is made up of 3D cubes called
voxels. 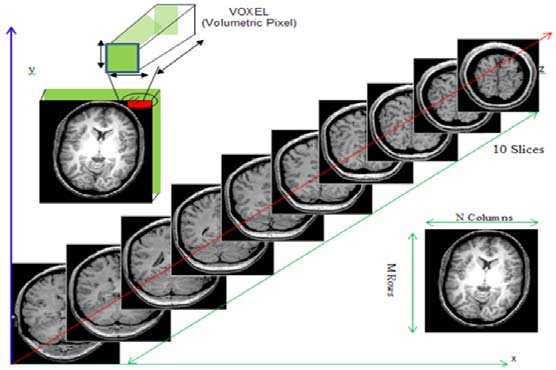
What about the size along each dimension (shape)?
The 3 numbers given here represent the number of values along a respective dimension (x,y,z). This brain was scanned in 57 slices with a resolution of 67 x 56 voxels per slice. That means there are:
57 * 67 * 56 = 213864
voxels in total!
Let’s see the type of data inside the array.
OUTPUT
dtype('float64')This tells us that each element in the array (or voxel) is a
floating-point number.
The data type of an image controls the range of possible intensities. As
the number of possible values increases, the amount of space the image
takes up in memory also increases.
OUTPUT
6.678363800048828
96.55541983246803For our data, the range of intensity values goes from 0 (black) to more positive digits (whiter).
How do we examine what value a particular voxel is? We can inspect the value of a voxel by selecting an index as follows:
data[x,y,z]
So for example we can inspect a voxel at coordinates (10,20,3) by doing the following:
OUTPUT
32.40787395834923NOTE: Python uses zero-based indexing. The first item in the array is item 0. The second item is item 1, the third is item 2, etc.
This yields a single value representing the intensity of the signal at a particular voxel! Next we’ll see how to not just pull one voxel but a slice or an array of voxels for visualization and analysis!
Working with image data
Slicing does exactly what it seems to imply. Giving our 3D volume, we pull out a 2D slice of our data.
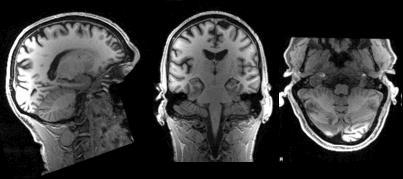 From left to right:
sagittal, coronal and axial slices.
From left to right:
sagittal, coronal and axial slices.
Let’s pull the 10th slice in the x axis.
This is similar to the indexing we did before to pull out a single
voxel. However, instead of providing a value for each axis, the
: indicates that we want to grab all values from
that particular axis.
Slicing MRI data
Now try selecting the 20th slice from the y axis.
Slicing MRI data (continued)
Finally, try grabbing the 3rd slice from the z axis
We’ve been slicing and dicing brain images but we have no idea what they look like! In the next section we’ll show you how you can visualize brain slices!
Visualizing the data
We previously inspected the signal intensity of the voxel at
coordinates (10,20,3). Let’s see what out data looks like when we slice
it at this location. We’ve already indexed the data at each x, y, and z
axis. Let’s use matplotlib.
PYTHON
import matplotlib.pyplot as plt
%matplotlib inline
slices = [x_slice, y_slice, z_slice]
fig, axes = plt.subplots(1, len(slices))
for i, slice in enumerate(slices):
axes[i].imshow(slice.T, cmap="gray", origin="lower")Now, we’re going to step away from discussing our data and talk about the final important attribute of a NIfTI.
3. Affine: tells the position of the image array data in a reference space
The final important piece of metadata associated with an image file is the affine matrix. Below is the affine matrix for our data.
OUTPUT
array([[ 2.75, 0. , 0. , -78. ],
[ 0. , 2.75, 0. , -91. ],
[ 0. , 0. , 2.75, -91. ],
[ 0. , 0. , 0. , 1. ]])To explain this concept, recall that we referred to coordinates in our data as (x,y,z) coordinates such that:
- x is the first dimension of
t1_data - y is the second dimension of
t1_data - z is the third dimension of
t1_data
Although this tells us how to access our data in terms of voxels in a 3D volume, it doesn’t tell us much about the actual dimensions in our data (centimetres, right or left, up or down, back or front). The affine matrix allows us to translate between voxel coordinates in (x,y,z) and world space coordinates in (left/right,bottom/top,back/front). An important thing to note is that in reality in which order you have:
- left/right
- bottom/top
- back/front
Depends on how you’ve constructed the affine matrix, but for the data we’re dealing with it always refers to:
- Right
- Anterior
- Superior
Applying the affine matrix (t1_affine) is done through
using a linear map (matrix multiplication) on voxel coordinates
(defined in t1_data).
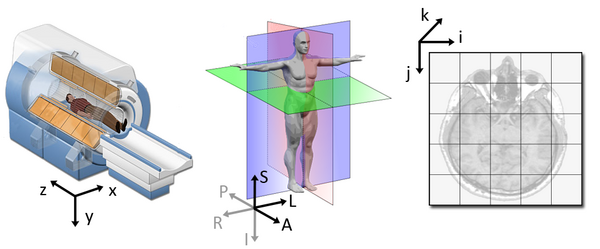
The concept of an affine matrix may seem confusing at first but an example might help gain an intuition:
Suppose we have two voxels located at the following coordinates: \((64, 100, 2)\)
And we wanted to know what the distances between these two voxels are in terms of real world distances (millimetres). This information cannot be derived from using voxel coordinates so we turn to the affine matrix.
Now, the affine matrix we’ll be using happens to be encoded in
RAS. That means once we apply the matrix our
coordinates are as follows:(Right, Anterior, Superior)
So increasing a coordinate value in the first dimension corresponds to moving to the right of the person being scanned.
Applying our affine matrix yields the following coordinates: \((10.25, 30.5, 9.2)\)
This means that:
- Voxel 1 is
90.23 - 10.25 = 79.98in the R axis. Positive values mean move right - Voxel 1 is
0.2 - 30.5 = -30.3in the A axis. Negative values mean move posterior - Voxel 1 is
2.15 - 9.2 = -7.05in the S axis. Negative values mean move inferior
This covers the basics of how NIfTI data and metadata are stored and organized in the context of Python. In the next segment we’ll talk a bit about an increasingly important component of MR data analysis - data organization. This is a key component to reproducible analysis and so we’ll spend a bit of time here.
- NIfTI image contain a header, which describes the contents, and the data.
- The position of the NIfTI data in space is determined by the affine matrix.
- NIfTI data is a multi-dimensional array of values.
Content from Data organization with BIDS
Last updated on 2025-02-26 | Edit this page
Overview
Questions
- What is BIDS?
- Why is it important to structure our datasets coherently?
- How can we convert raw datasets into BIDS-compatible datasets?
Objectives
- Become faimiliar with BIDS
- Convert raw datasets to BIDS-compatible datasets
Brain Imaging Data Structure (BIDS) is a simple and intuitive way to organize and describe your neuroimaging and behavioural data. Neuroimaging experiments result in complicated data that can be arranged in several different ways. BIDS tackles this problem by suggesting a new standard (based on consensus from multiple researchers across the world) for the arrangement of neuroimaging datasets.
Using the same structure for all your studies will allow you to easily reuse all your scripts between studies. Additionally, sharing code with other researchers will be much easier.
Why convert the community to BIDS?
Share data:
- A common file structure for both computers and humans to understand
- Using file formats/conventions that are most common in neuroimaging
- Show completeness of the data right away
Content from Exploring open MRI datasets
Last updated on 2024-02-28 | Edit this page
Overview
Questions
- How does standardizing neuroimaging data ease the data exploration process
Objectives
- Understand the BIDS format
- Use PyBIDS in to easily explore a BIDS dataset
Tutorial Dataset
In this episode, we will be using a subset of a publicly available dataset, ds000030, from openneuro.org. All of the datasets on OpenNeuro are already structured according to BIDS.
Downloading Data
DataLad
DataLad installs the data - which for a dataset means
that we get the “small” data (i.e. the text files) and the download
instructions for the larger files. We can now navigate the dataset like
its a part of our file system and plan our analysis.
First, navigate to the folder where you’d like to download the dataset.
Let’s take a look at the participants.tsv file to see
what the demographics for this dataset look like.
PYTHON
import pandas as pd
participant_metadata = pd.read_csv('../data/ds000030/participants.tsv', sep='\t')
participant_metadataOUTPUT:
| participant_id | diagnosis | age | gender | bart | bht | dwi | pamenc | pamret | rest | scap | stopsignal | T1w | taskswitch | ScannerSerialNumber | ghost_NoGhost | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | sub-10159 | CONTROL | 30 | F | 1.0 | NaN | 1.0 | NaN | NaN | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 35343.0 | No_ghost |
| 1 | sub-10171 | CONTROL | 24 | M | 1.0 | 1.0 | 1.0 | NaN | NaN | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 35343.0 | No_ghost |
| 2 | sub-10189 | CONTROL | 49 | M | 1.0 | NaN | 1.0 | NaN | NaN | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 35343.0 | No_ghost |
| 3 | sub-10193 | CONTROL | 40 | M | 1.0 | NaN | 1.0 | NaN | NaN | NaN | NaN | NaN | 1.0 | NaN | 35343.0 | No_ghost |
| 4 | sub-10206 | CONTROL | 21 | M | 1.0 | NaN | 1.0 | NaN | NaN | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 35343.0 | No_ghost |
From this table we can easily view unique diagnostic groups:
OUTPUT
array(['CONTROL', 'SCHZ', 'BIPOLAR', 'ADHD'], dtype=object)Imagine we’d like to work with participants that are either CONTROL
or SCHZ (diagnosis) and have both a T1w
(T1w == 1) and rest (rest == 1) scan. Also,
you’ll notice some of the T1w scans included in this dataset have a
ghosting artifact. We’ll need to filter these out as well
(ghost_NoGhost == 'No_ghost').
We’ll filter this data out like so:
PYTHON
participant_metadata = participant_metadata[(participant_metadata.diagnosis.isin(['CONTROL', 'SCHZ'])) &
(participant_metadata.T1w == 1) &
(participant_metadata.rest == 1) &
(participant_metadata.ghost_NoGhost == 'No_ghost')]
participant_metadata['diagnosis'].unique()OUTPUT
array(['CONTROL', 'SCHZ'], dtype=object)From this we have a list of participants corresponding to a list of
participants who are of interest in our analysis. We can then use this
list in order to download participant from software such as
aws or datalad. In fact, this is exactly how
we set up a list of participants to download for the fMRI workshop!
Since we’ve already downloaded the dataset, we can now explore the
structure using PyBIDS, a Python API for querying, summarizing and
manipulating the BIDS folder structure.
Getting and dropping data
datalad get datalad drop
The pybids layout object indexes the BIDS folder. Indexing can take a
really long time, especially if you have several subjects, modalities,
scan types, etc. pybids has an option to save the indexed
results to a SQLite database. This database can then be re-used the next
time you want to query the same BIDS folder.
PYTHON
layout.save("~/Desktop/dc-mri/data/ds000030/.db")
layout = BIDSLayout("~/Desktop/dc-mri/data/ds000030", database_path="~/Desktop/dc-mri/data/ds000030/.db")The pybids layout object also lets you query your BIDS dataset
according to a number of parameters by using a get_*()
method. We can get a list of the subjects we’ve downloaded from the
dataset.
OUTPUT
['10171',
'10292',
'10365',
'10438',
'10565',
'10788',
'11106',
'11108',
'11122',
...
'50083']We can also pull a list of imaging modalities in the dataset:
OUTPUT
['anat', 'func']As well as tasks and more!:
PYTHON
#Task fMRI
print(layout.get_tasks())
#Data types (bold, brainmask, confounds, smoothwm, probtissue, warp...)
print(layout.get_types())OUTPUT
['rest']
['bold',
'brainmask',
'confounds',
'description',
'dtissue',
'fsaverage5',
'inflated',
'midthickness',
'participants',
'pial',
'preproc',
'probtissue',
'smoothwm',
'warp']In addition we can specify sub-types of each BIDS category:
OUTPUT
['brainmask', 'confounds', 'fsaverage5', 'preproc']We can use this functionality to pull all our fMRI NIfTI files:
OUTPUT
TO FILLFinally, we can convert the data stored in bids.layout
into a pandas.DataFrame :
OUTPUT:
| path | modality | subject | task | type | |
|---|---|---|---|---|---|
| 0 | ~/Desktop/dc-mri/data… | NaN | NaN | rest | bold |
| 1 | ~/Desktop/dc-mri/data… | NaN | NaN | NaN | participants |
| 2 | ~/Desktop/dc-mri/data… | NaN | NaN | NaN | NaN |
| 3 | ~/Desktop/dc-mri/data… | anat | 10565 | NaN | brainmask |
| 4 | ~/Desktop/dc-mri/data… | anat | 10565 | NaN | probtissue |
- Public neuroimaging BIDS-compatible data repositories allow for pulling data easily.
- PyBIDS is a Python-based tool that allows for easy exploration of BIDS-formatted neuroimaging data.
Content from BIDS derivatives
Last updated on 2025-02-26 | Edit this page
Overview
Questions
- What are BIDS Apps?
- What are containers?
Objectives
- Learn about neuroimaging applications using BIDS data structures
- Use neuroimaging applications through containers
BIDS Apps are containerized applications that run on BIDS data structures.
Some examples include:
- mriqc
- fmriprep
- freesurfer
- ciftify
- SPM
- MRtrix3_connectome
They rely on 2 technologies for container computing:
-
Docker
- for building, hosting, and running containers on local hardware (Windows, Mac OS, Linux) or in the cloud
-
Singularity
- for running containers on high performance compute clusters
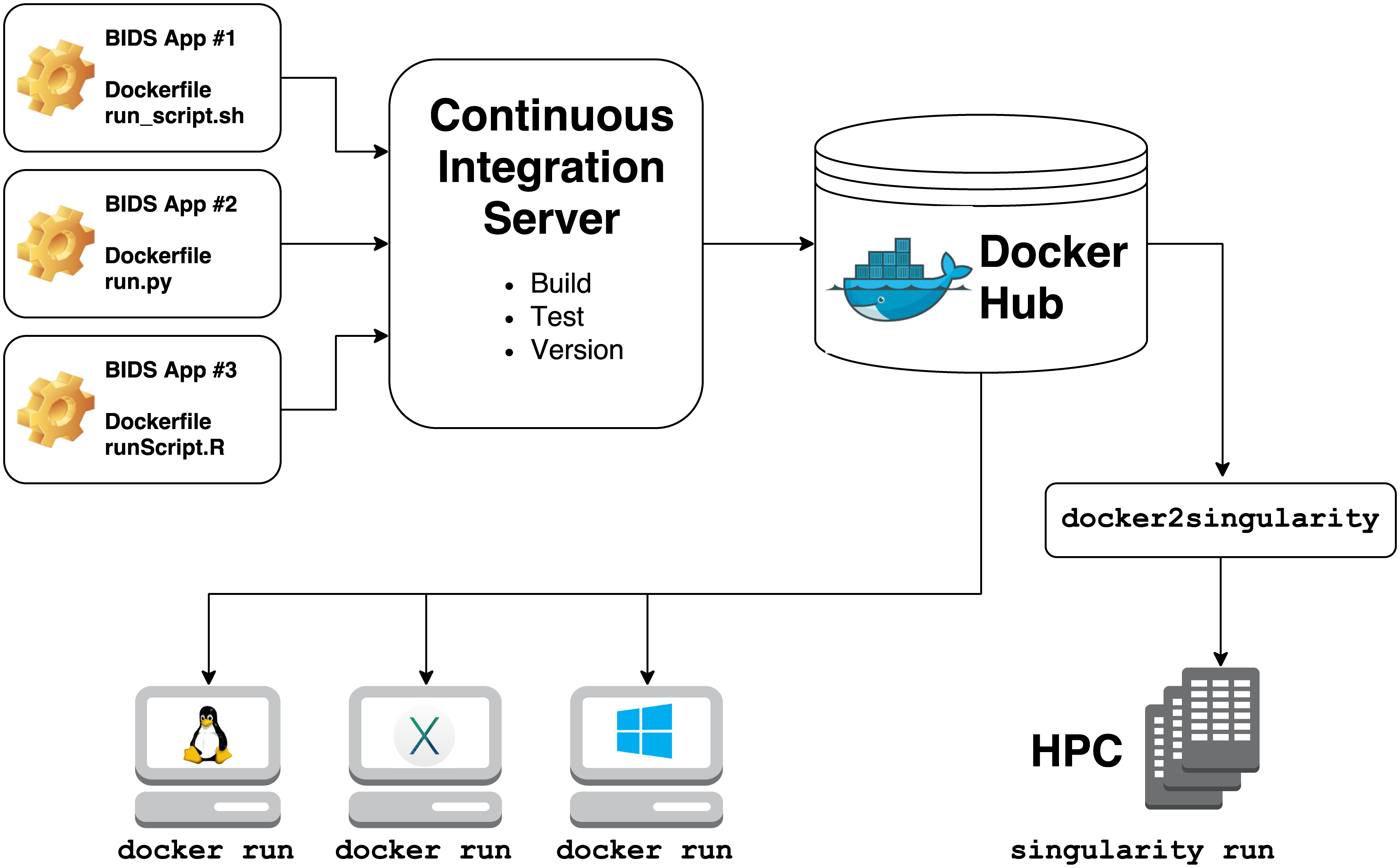
Building a singularity container is as easy as:
To run the container:
BASH
singularity run --cleanenv \
-B bids_folder:/data \
mriqc-0.16.1.simg \
/data /data/derivatives participant- BIDS Apps are containerized applications that run on BIDS-compatible datasets.