Why use a Cluster?
Overview
Teaching: 15 min
Exercises: 5 minQuestions
Why would I be interested in High Performance Computing (HPC)?
What can I expect to learn from this course?
Objectives
Describe what an HPC system is
Identify how an HPC system could benefit you.
Frequently, research problems that use computing can outgrow the capabilities of the desktop or laptop computer where they started:
- A statistics student wants to cross-validate a model. This involves running the model 1000 times – but each run takes an hour. Running the model on a laptop will take over a month! In this research problem, final results are calculated after all 1000 models have run, but typically only one model is run at a time (in serial) on the laptop. Since each of the 1000 runs is independent of all others, and given enough computers, it’s theoretically possible to run them all at once (in parallel).
- A genomics researcher has been using small datasets of sequence data, but soon will be receiving a new type of sequencing data that is 10 times as large. It’s already challenging to open the datasets on a computer – analyzing these larger datasets will probably crash it. In this research problem, the calculations required might be impossible to parallelize, but a computer with more memory would be required to analyze the much larger future data set.
- An engineer is using a fluid dynamics package that has an option to run in parallel. So far, this option was not used on a desktop. In going from 2D to 3D simulations, the simulation time has more than tripled. It might be useful to take advantage of that option or feature. In this research problem, the calculations in each region of the simulation are largely independent of calculations in other regions of the simulation. It’s possible to run each region’s calculations simultaneously (in parallel), communicate selected results to adjacent regions as needed, and repeat the calculations to converge on a final set of results. In moving from a 2D to a 3D model, both the amount of data and the amount of calculations increases greatly, and it’s theoretically possible to distribute the calculations across multiple computers communicating over a shared network.
In all these cases, access to more (and larger) computers is needed. Those computers should be usable at the same time, solving many researchers’ problems in parallel.
Jargon Busting Presentation
Open the HPC Jargon Buster
in a new tab. To present the content, press C to open a clone in a
separate window, then press P to toggle presentation mode.
I’ve Never Used a Server, Have I?
Take a minute and think about which of your daily interactions with a computer may require a remote server or even cluster to provide you with results.
Some Ideas
- Checking email: your computer (possibly in your pocket) contacts a remote machine, authenticates, and downloads a list of new messages; it also uploads changes to message status, such as whether you read, marked as junk, or deleted the message. Since yours is not the only account, the mail server is probably one of many in a data center.
- Searching for a phrase online involves comparing your search term against a massive database of all known sites, looking for matches. This “query” operation can be straightforward, but building that database is a monumental task! Servers are involved at every step.
- Searching for directions on a mapping website involves connecting your (A) starting and (B) end points by traversing a graph in search of the “shortest” path by distance, time, expense, or another metric. Converting a map into the right form is relatively simple, but calculating all the possible routes between A and B is expensive.
Checking email could be serial: your machine connects to one server and exchanges data. Searching by querying the database for your search term (or endpoints) could also be serial, in that one machine receives your query and returns the result. However, assembling and storing the full database is far beyond the capability of any one machine. Therefore, these functions are served in parallel by a large, “hyperscale” collection of servers working together.
Key Points
High Performance Computing (HPC) typically involves connecting to very large computing systems elsewhere in the world.
These other systems can be used to do work that would either be impossible or much slower on smaller systems.
HPC resources are shared by multiple users.
The standard method of interacting with such systems is via a command line interface.
Connecting to a remote HPC system
Overview
Teaching: 25 min
Exercises: 10 minQuestions
How do I log in to a remote HPC system?
Objectives
Configure secure access to a remote HPC system.
Connect to a remote HPC system.
Secure Connections
The first step in using a cluster is to establish a connection from our laptop to the cluster. When we are sitting at a computer (or standing, or holding it in our hands or on our wrists), we have come to expect a visual display with icons, widgets, and perhaps some windows or applications: a graphical user interface, or GUI. Since computer clusters are remote resources that we connect to over slow or intermittent interfaces (WiFi and VPNs especially), it is more practical to use a command-line interface, or CLI, to send commands as plain-text. If a command returns output, it is printed as plain text as well. The commands we run today will not open a window to show graphical results.
If you have ever opened the Windows Command Prompt or macOS Terminal, you have seen a CLI. If you have already taken The Carpentries’ courses on the UNIX Shell or Version Control, you have used the CLI on your local machine extensively. The only leap to be made here is to open a CLI on a remote machine, while taking some precautions so that other folks on the network can’t see (or change) the commands you’re running or the results the remote machine sends back. We will use the Secure SHell protocol (or SSH) to open an encrypted network connection between two machines, allowing you to send & receive text and data without having to worry about prying eyes.
SSH clients are usually command-line tools, where you provide the remote
machine address as the only required argument. If your username on the remote
system differs from what you use locally, you must provide that as well. If
your SSH client has a graphical front-end, such as PuTTY or MobaXterm, you will
set these arguments before clicking “connect.” From the terminal, you’ll write
something like ssh userName@hostname, where the argument is just like an
email address: the “@” symbol is used to separate the personal ID from the
address of the remote machine.
When logging in to a laptop, tablet, or other personal device, a username, password, or pattern are normally required to prevent unauthorized access. In these situations, the likelihood of somebody else intercepting your password is low, since logging your keystrokes requires a malicious exploit or physical access. For systems like login1 running an SSH server, anybody on the network can log in, or try to. Since usernames are often public or easy to guess, your password is often the weakest link in the security chain. Many clusters therefore forbid password-based login, requiring instead that you generate and configure a public-private key pair with a much stronger password. Even if your cluster does not require it, the next section will guide you through the use of SSH keys and an SSH agent to both strengthen your security and make it more convenient to log in to remote systems.
Better Security With SSH Keys
The Lesson Setup provides instructions for installing a shell application with SSH. If you have not done so already, please open that shell application with a Unix-like command line interface to your system.
SSH keys are an alternative method for authentication to obtain access to remote computing systems. They can also be used for authentication when transferring files or for accessing remote version control systems (such as GitHub). In this section you will create a pair of SSH keys:
- a private key which you keep on your own computer, and
- a public key which can be placed on any remote system you will access.
Private keys are your secure digital passport
A private key that is visible to anyone but you should be considered compromised, and must be destroyed. This includes having improper permissions on the directory it (or a copy) is stored in, traversing any network that is not secure (encrypted), attachment on unencrypted email, and even displaying the key on your terminal window.
Protect this key as if it unlocks your front door. In many ways, it does.
Regardless of the software or operating system you use, please choose a strong password or passphrase to act as another layer of protection for your private SSH key.
Considerations for SSH Key Passwords
When prompted, enter a strong password that you will remember. There are two common approaches to this:
- Create a memorable passphrase with some punctuation and number-for-letter substitutions, 32 characters or longer. Street addresses work well; just be careful of social engineering or public records attacks.
- Use a password manager and its built-in password generator with all character classes, 25 characters or longer. KeePass and BitWarden are two good options.
- Nothing is less secure than a private key with no password. If you skipped password entry by accident, go back and generate a new key pair with a strong password.
SSH Keys on Linux, Mac, MobaXterm, and Windows Subsystem for Linux
Once you have opened a terminal, check for existing SSH keys and filenames since existing SSH keys are overwritten.
[you@laptop:~]$ ls ~/.ssh/
If ~/.ssh/id_ed25519 already exists, you will need to specify
a different name for the new key-pair.
Generate a new public-private key pair using the following command, which will
produce a stronger key than the ssh-keygen default by invoking these flags:
-a(default is 16): number of rounds of passphrase derivation; increase to slow down brute force attacks.-t(default is rsa): specify the “type” or cryptographic algorithm.ed25519specifies EdDSA with a 256-bit key; it is faster than RSA with a comparable strength.-f(default is /home/user/.ssh/id_algorithm): filename to store your private key. The public key filename will be identical, with a.pubextension added.
[you@laptop:~]$ ssh-keygen -a 100 -f ~/.ssh/id_ed25519 -t ed25519
When prompted, enter a strong password with the above considerations in mind. Note that the terminal will not appear to change while you type the password: this is deliberate, for your security. You will be prompted to type it again, so don’t worry too much about typos.
Take a look in ~/.ssh (use ls ~/.ssh). You should see two new files:
- your private key (
~/.ssh/id_ed25519): do not share with anyone! - the shareable public key (
~/.ssh/id_ed25519.pub): if a system administrator asks for a key, this is the one to send. It is also safe to upload to websites such as GitHub: it is meant to be seen.
Use RSA for Older Systems
If key generation failed because ed25519 is not available, try using the older (but still strong and trustworthy) RSA cryptosystem. Again, first check for an existing key:
[you@laptop:~]$ ls ~/.ssh/If
~/.ssh/id_rsaalready exists, you will need to specify choose a different name for the new key-pair. Generate it as above, with the following extra flags:
-bsets the number of bits in the key. The default is 2048. EdDSA uses a fixed key length, so this flag would have no effect.-o(no default): use the OpenSSH key format, rather than PEM.[you@laptop:~]$ ssh-keygen -a 100 -b 4096 -f ~/.ssh/id_rsa -o -t rsaWhen prompted, enter a strong password with the above considerations in mind.
Take a look in
~/.ssh(usels ~/.ssh). You should see two new files:
- your private key (
~/.ssh/id_rsa): do not share with anyone!- the shareable public key (
~/.ssh/id_rsa.pub): if a system administrator asks for a key, this is the one to send. It is also safe to upload to websites such as GitHub: it is meant to be seen.
SSH Keys on PuTTY
If you are using PuTTY on Windows, download and use puttygen to generate the
key pair. See the PuTTY documentation for details.
- Select
EdDSAas the key type. - Select
255as the key size or strength. - Click on the “Generate” button.
- You do not need to enter a comment.
- When prompted, enter a strong password with the above considerations in mind.
- Save the keys in a folder no other users of the system can read.
Take a look in the folder you specified. You should see two new files:
- your private key (
id_ed25519): do not share with anyone! - the shareable public key (
id_ed25519.pub): if a system administrator asks for a key, this is the one to send. It is also safe to upload to websites such as GitHub: it is meant to be seen.
SSH Agent for Easier Key Handling
An SSH key is only as strong as the password used to unlock it, but on the other hand, typing out a complex password every time you connect to a machine is tedious and gets old very fast. This is where the SSH Agent comes in.
Using an SSH Agent, you can type your password for the private key once, then have the Agent remember it for some number of hours or until you log off. Unless some nefarious actor has physical access to your machine, this keeps the password safe, and removes the tedium of entering the password multiple times.
Just remember your password, because once it expires in the Agent, you have to type it in again.
SSH Agents on Linux, macOS, and Windows
Open your terminal application and check if an agent is running:
[you@laptop:~]$ ssh-add -l
-
If you get an error like this one,
Error connecting to agent: No such file or directory… then you need to launch the agent as follows:
[you@laptop:~]$ eval $(ssh-agent)What’s in a
$(...)?The syntax of this SSH Agent command is unusual, based on what we’ve seen in the UNIX Shell lesson. This is because the
ssh-agentcommand creates opens a connection that only you have access to, and prints a series of shell commands that can be used to reach it – but does not execute them![you@laptop:~]$ ssh-agentSSH_AUTH_SOCK=/tmp/ssh-Zvvga2Y8kQZN/agent.131521; export SSH_AUTH_SOCK; SSH_AGENT_PID=131522; export SSH_AGENT_PID; echo Agent pid 131522;The
evalcommand interprets this text output as commands and allows you to access the SSH Agent connection you just created.You could run each line of the
ssh-agentoutput yourself, and achieve the same result. Usingevaljust makes this easier. -
Otherwise, your agent is already running: don’t mess with it.
Add your key to the agent, with session expiration after 8 hours:
[you@laptop:~]$ ssh-add -t 8h ~/.ssh/id_ed25519
Enter passphrase for .ssh/id_ed25519:
Identity added: .ssh/id_ed25519
Lifetime set to 86400 seconds
For the duration (8 hours), whenever you use that key, the SSH Agent will provide the key on your behalf without you having to type a single keystroke.
SSH Agent on PuTTY
If you are using PuTTY on Windows, download and use pageant as the SSH agent.
See the PuTTY documentation.
Transfer Your Public Key
Visit https://mokey.cluster.hpc-carpentry.org to upload your SSH
public key. (Remember, it’s the one ending in .pub!)
Log In to the Cluster
Go ahead and open your terminal or graphical SSH client, then log in to the
cluster. Replace yourUsername with your username or the one
supplied by the instructors.
[you@laptop:~]$ ssh yourUsername@cluster.hpc-carpentry.org
You may be asked for your password. Watch out: the characters you type after
the password prompt are not displayed on the screen. Normal output will resume
once you press Enter.
You may have noticed that the prompt changed when you logged into the remote
system using the terminal (if you logged in using PuTTY this will not apply
because it does not offer a local terminal). This change is important because
it can help you distinguish on which system the commands you type will be run
when you pass them into the terminal. This change is also a small complication
that we will need to navigate throughout the workshop. Exactly what is displayed
as the prompt (which conventionally ends in $) in the terminal when it is
connected to the local system and the remote system will typically be different
for every user. We still need to indicate which system we are entering commands
on though so we will adopt the following convention:
[you@laptop:~]$when the command is to be entered on a terminal connected to your local computer[yourUsername@login1 ~]$when the command is to be entered on a terminal connected to the remote system$when it really doesn’t matter which system the terminal is connected to.
Looking Around Your Remote Home
Very often, many users are tempted to think of a high-performance computing
installation as one giant, magical machine. Sometimes, people will assume that
the computer they’ve logged onto is the entire computing cluster. So what’s
really happening? What computer have we logged on to? The name of the current
computer we are logged onto can be checked with the hostname command. (You
may also notice that the current hostname is also part of our prompt!)
[yourUsername@login1 ~]$ hostname
login1
So, we’re definitely on the remote machine. Next, let’s find out where we are
by running pwd to print the working directory.
[yourUsername@login1 ~]$ pwd
/home/yourUsername
Great, we know where we are! Let’s see what’s in our current directory:
[yourUsername@login1 ~]$ ls
id_ed25519.pub
The system administrators may have configured your home directory with some helpful files, folders, and links (shortcuts) to space reserved for you on other filesystems. If they did not, your home directory may appear empty. To double-check, include hidden files in your directory listing:
[yourUsername@login1 ~]$ ls -a
. .bashrc id_ed25519.pub
.. .ssh
In the first column, . is a reference to the current directory and .. a
reference to its parent (/home). You may or may not see
the other files, or files like them: .bashrc is a shell configuration file,
which you can edit with your preferences; and .ssh is a directory storing SSH
keys and a record of authorized connections.
Key Points
An HPC system is a set of networked machines.
HPC systems typically provide login nodes and a set of worker nodes.
The resources found on independent (worker) nodes can vary in volume and type (amount of RAM, processor architecture, availability of network mounted filesystems, etc.).
Files saved on one node are available on all nodes.
Exploring Remote Resources
Overview
Teaching: 25 min
Exercises: 10 minQuestions
How does my local computer compare to the remote systems?
How does the login node compare to the compute nodes?
Are all compute nodes alike?
Objectives
Survey system resources using
nproc,free, and the queuing systemCompare & contrast resources on the local machine, login node, and worker nodes
Learn about the various filesystems on the cluster using
dfFind out
whoelse is logged inAssess the number of idle and occupied nodes
Look Around the Remote System
If you have not already connected to HPC Carpentry’s Cloud Cluster, please do so now:
[you@laptop:~]$ ssh yourUsername@cluster.hpc-carpentry.org
Take a look at your home directory on the remote system:
[yourUsername@login1 ~]$ ls
What’s different between your machine and the remote?
Open a second terminal window on your local computer and run the
lscommand (without logging in to HPC Carpentry’s Cloud Cluster). What differences do you see?Solution
You would likely see something more like this:
[you@laptop:~]$ lsApplications Documents Library Music Public Desktop Downloads Movies PicturesThe remote computer’s home directory shares almost nothing in common with the local computer: they are completely separate systems!
Most high-performance computing systems run the Linux operating system, which
is built around the UNIX Filesystem Hierarchy Standard. Instead of
having a separate root for each hard drive or storage medium, all files and
devices are anchored to the “root” directory, which is /:
[yourUsername@login1 ~]$ ls /
bin etc lib64 proc sbin sys var
boot home mnt root scratch tmp working
dev lib opt run srv usr
The “home” directory is the one where we generally want to keep all of our files. Other folders on a UNIX OS contain system files and change as you install new software or upgrade your OS.
Using HPC filesystems
On HPC systems, you have a number of places where you can store your files. These differ in both the amount of space allocated and whether or not they are backed up.
- Home – often a network filesystem, data stored here is available throughout the HPC system, and often backed up periodically. Files stored here are typically slower to access, the data is actually stored on another computer and is being transmitted and made available over the network!
- Scratch – typically faster than the networked Home directory, but not usually backed up, and should not be used for long term storage.
- Work – sometimes provided as an alternative to Scratch space, Work is a fast file system accessed over the network. Typically, this will have higher performance than your home directory, but lower performance than Scratch; it may not be backed up. It differs from Scratch space in that files in a work file system are not automatically deleted for you: you must manage the space yourself.
Nodes
Individual computers that compose a cluster are typically called nodes (although you will also hear people call them servers, computers and machines). On a cluster, there are different types of nodes for different types of tasks. The node where you are right now is called the login node, head node, landing pad, or submit node. A login node serves as an access point to the cluster.
As a gateway, the login node should not be used for time-consuming or resource-intensive tasks. You should be alert to this, and check with your site’s operators or documentation for details of what is and isn’t allowed. It is well suited for uploading and downloading files, setting up software, and running tests. Generally speaking, in these lessons, we will avoid running jobs on the login node.
Who else is logged in to the login node?
[yourUsername@login1 ~]$ who
This may show only your user ID, but there are likely several other people (including fellow learners) connected right now.
Dedicated Transfer Nodes
If you want to transfer larger amounts of data to or from the cluster, some systems offer dedicated nodes for data transfers only. The motivation for this lies in the fact that larger data transfers should not obstruct operation of the login node for anybody else. Check with your cluster’s documentation or its support team if such a transfer node is available. As a rule of thumb, consider all transfers of a volume larger than 500 MB to 1 GB as large. But these numbers change, e.g., depending on the network connection of yourself and of your cluster or other factors.
The real work on a cluster gets done by the compute (or worker) nodes. compute nodes come in many shapes and sizes, but generally are dedicated to long or hard tasks that require a lot of computational resources.
All interaction with the compute nodes is handled by a specialized piece of software called a scheduler (the scheduler used in this lesson is called Slurm). We’ll learn more about how to use the scheduler to submit jobs next, but for now, it can also tell us more information about the compute nodes.
For example, we can view all of the compute nodes by running the command
sinfo.
[yourUsername@login1 ~]$ sinfo
PARTITION AVAIL TIMELIMIT NODES STATE NODELIST
cpubase_bycore_b1* up infinite 4 idle node[1-2],smnode[1-2]
node up infinite 2 idle node[1-2]
smnode up infinite 2 idle smnode[1-2]
A lot of the nodes are busy running work for other users: we are not alone here!
There are also specialized machines used for managing disk storage, user authentication, and other infrastructure-related tasks. Although we do not typically logon to or interact with these machines directly, they enable a number of key features like ensuring our user account and files are available throughout the HPC system.
What’s in a Node?
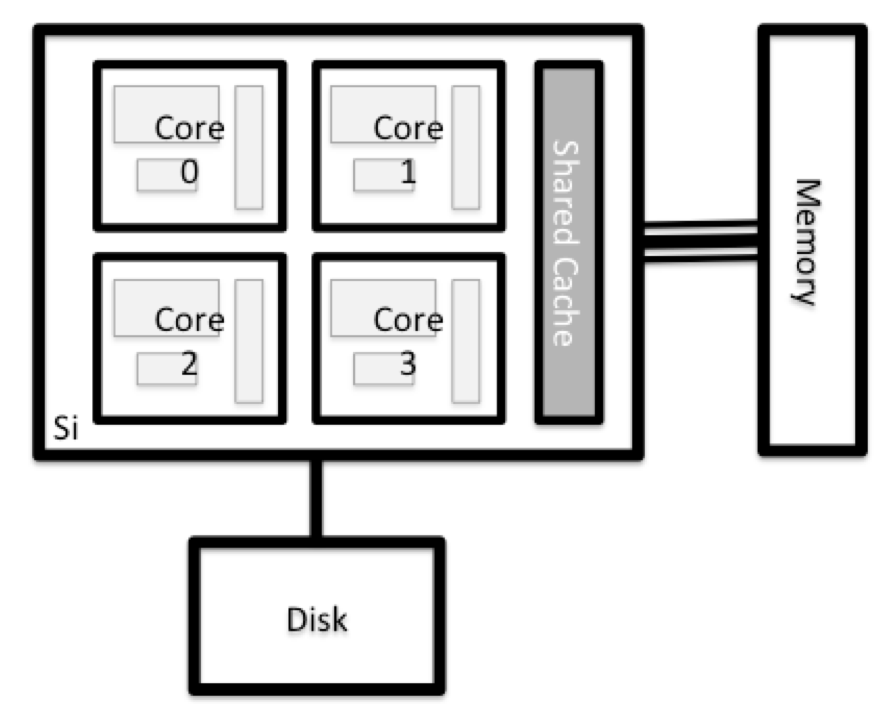
All of the nodes in an HPC system have the same components as your own laptop or desktop: CPUs (sometimes also called processors or cores), memory (or RAM), and disk space. CPUs are a computer’s tool for actually running programs and calculations. Information about a current task is stored in the computer’s memory. Disk refers to all storage that can be accessed like a file system. This is generally storage that can hold data permanently, i.e. data is still there even if the computer has been restarted. While this storage can be local (a hard drive installed inside of it), it is more common for nodes to connect to a shared, remote fileserver or cluster of servers.
Explore Your Computer
Try to find out the number of CPUs and amount of memory available on your personal computer.
Note that, if you’re logged in to the remote computer cluster, you need to log out first. To do so, type
Ctrl+dorexit:[yourUsername@login1 ~]$ exit [you@laptop:~]$Solution
There are several ways to do this. Most operating systems have a graphical system monitor, like the Windows Task Manager. More detailed information can be found on the command line:
- Run system utilities
[you@laptop:~]$ nproc --all [you@laptop:~]$ free -m- Read from
/proc[you@laptop:~]$ cat /proc/cpuinfo [you@laptop:~]$ cat /proc/meminfo- Run system monitor
[you@laptop:~]$ htop
Explore the Login Node
Now compare the resources of your computer with those of the login node.
Solution
[you@laptop:~]$ ssh yourUsername@cluster.hpc-carpentry.org [yourUsername@login1 ~]$ nproc --all [yourUsername@login1 ~]$ free -mYou can get more information about the processors using
lscpu, and a lot of detail about the memory by reading the file/proc/meminfo:[yourUsername@login1 ~]$ less /proc/meminfoYou can also explore the available filesystems using
dfto show disk free space. The-hflag renders the sizes in a human-friendly format, i.e., GB instead of B. The type flag-Tshows what kind of filesystem each resource is.[yourUsername@login1 ~]$ df -ThDifferent results from
df
- The local filesystems (ext, tmp, xfs, zfs) will depend on whether you’re on the same login node (or compute node, later on).
- Networked filesystems (beegfs, cifs, gpfs, nfs, pvfs) will be similar – but may include yourUsername, depending on how it is mounted.
Shared Filesystems
This is an important point to remember: files saved on one node (computer) are often available everywhere on the cluster!
Explore a Worker Node
Finally, let’s look at the resources available on the worker nodes where your jobs will actually run. Try running this command to see the name, CPUs and memory available on the worker nodes:
[yourUsername@login1 ~]$ sinfo -o "%n %c %m" | column -t
Compare Your Computer, the Login Node and the Compute Node
Compare your laptop’s number of processors and memory with the numbers you see on the cluster login node and compute node. What implications do you think the differences might have on running your research work on the different systems and nodes?
Solution
Compute nodes are usually built with processors that have higher core-counts than the login node or personal computers in order to support highly parallel tasks. Compute nodes usually also have substantially more memory (RAM) installed than a personal computer. More cores tends to help jobs that depend on some work that is easy to perform in parallel, and more, faster memory is key for large or complex numerical tasks.
Differences Between Nodes
Many HPC clusters have a variety of nodes optimized for particular workloads. Some nodes may have larger amount of memory, or specialized resources such as Graphics Processing Units (GPUs or “video cards”).
With all of this in mind, we will now cover how to talk to the cluster’s scheduler, and use it to start running our scripts and programs!
Key Points
An HPC system is a set of networked machines.
HPC systems typically provide login nodes and a set of compute nodes.
The resources found on independent (worker) nodes can vary in volume and type (amount of RAM, processor architecture, availability of network mounted filesystems, etc.).
Files saved on shared storage are available on all nodes.
The login node is a shared machine: be considerate of other users.
Scheduler Fundamentals
Overview
Teaching: 45 min
Exercises: 30 minQuestions
What is a scheduler and why does a cluster need one?
How do I launch a program to run on a compute node in the cluster?
How do I capture the output of a program that is run on a node in the cluster?
Objectives
Submit a simple script to the cluster.
Monitor the execution of jobs using command line tools.
Inspect the output and error files of your jobs.
Find the right place to put large datasets on the cluster.
Job Scheduler
An HPC system might have thousands of nodes and thousands of users. How do we decide who gets what and when? How do we ensure that a task is run with the resources it needs? This job is handled by a special piece of software called the scheduler. On an HPC system, the scheduler manages which jobs run where and when.
The following illustration compares these tasks of a job scheduler to a waiter in a restaurant. If you can relate to an instance where you had to wait for a while in a queue to get in to a popular restaurant, then you may now understand why sometimes your job do not start instantly as in your laptop.
The scheduler used in this lesson is Slurm. Although Slurm is not used everywhere, running jobs is quite similar regardless of what software is being used. The exact syntax might change, but the concepts remain the same.
Running a Batch Job
The most basic use of the scheduler is to run a command non-interactively. Any command (or series of commands) that you want to run on the cluster is called a job, and the process of using a scheduler to run the job is called batch job submission.
In this case, the job we want to run is a shell script – essentially a text file containing a list of UNIX commands to be executed in a sequential manner. Our shell script will have three parts:
- On the very first line, add
#!/bin/bash. The#!(pronounced “hash-bang” or “shebang”) tells the computer what program is meant to process the contents of this file. In this case, we are telling it that the commands that follow are written for the command-line shell (what we’ve been doing everything in so far). - Anywhere below the first line, we’ll add an
echocommand with a friendly greeting. When run, the shell script will print whatever comes afterechoin the terminal.echo -nwill print everything that follows, without ending the line by printing the new-line character.
- On the last line, we’ll invoke the
hostnamecommand, which will print the name of the machine the script is run on.
[yourUsername@login1 ~]$ nano example-job.sh
#!/bin/bash
echo -n "This script is running on "
hostname
Creating Our Test Job
Run the script. Does it execute on the cluster or just our login node?
Solution
[yourUsername@login1 ~]$ bash example-job.shThis script is running on login1
This script ran on the login node, but we want to take advantage of
the compute nodes: we need the scheduler to queue up example-job.sh
to run on a compute node.
To submit this task to the scheduler, we use the
sbatch command.
This creates a job which will run the script when dispatched to
a compute node which the queuing system has identified as being
available to perform the work.
[yourUsername@login1 ~]$ sbatch example-job.sh
Submitted batch job 7
And that’s all we need to do to submit a job. Our work is done – now the
scheduler takes over and tries to run the job for us. While the job is waiting
to run, it goes into a list of jobs called the queue. To check on our job’s
status, we check the queue using the command
squeue -u yourUsername.
[yourUsername@login1 ~]$ squeue -u yourUsername
JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON)
9 cpubase_b example- user01 R 0:05 1 node1
We can see all the details of our job, most importantly that it is in the R
or RUNNING state. Sometimes our jobs might need to wait in a queue
(PENDING) or have an error (E).
Where’s the Output?
On the login node, this script printed output to the terminal – but now, when
squeueshows the job has finished, nothing was printed to the terminal.Cluster job output is typically redirected to a file in the directory you launched it from. Use
lsto find andcatto read the file.
Customising a Job
The job we just ran used all of the scheduler’s default options. In a real-world scenario, that’s probably not what we want. The default options represent a reasonable minimum. Chances are, we will need more cores, more memory, more time, among other special considerations. To get access to these resources we must customize our job script.
Comments in UNIX shell scripts (denoted by #) are typically ignored, but
there are exceptions. For instance the special #! comment at the beginning of
scripts specifies what program should be used to run it (you’ll typically see
#!/usr/bin/env bash). Schedulers like Slurm also
have a special comment used to denote special scheduler-specific options.
Though these comments differ from scheduler to scheduler,
Slurm’s special comment is #SBATCH. Anything
following the #SBATCH comment is interpreted as an
instruction to the scheduler.
Let’s illustrate this by example. By default, a job’s name is the name of the
script, but the -J option can be used to change the
name of a job. Add an option to the script:
[yourUsername@login1 ~]$ cat example-job.sh
#!/bin/bash
#SBATCH -J hello-world
echo -n "This script is running on "
hostname
Submit the job and monitor its status:
[yourUsername@login1 ~]$ sbatch example-job.sh
[yourUsername@login1 ~]$ squeue -u yourUsername
JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON)
10 cpubase_b hello-wo user01 R 0:02 1 node1
Fantastic, we’ve successfully changed the name of our job!
Resource Requests
What about more important changes, such as the number of cores and memory for our jobs? One thing that is absolutely critical when working on an HPC system is specifying the resources required to run a job. This allows the scheduler to find the right time and place to schedule our job. If you do not specify requirements (such as the amount of time you need), you will likely be stuck with your site’s default resources, which is probably not what you want.
The following are several key resource requests:
-
--ntasks=<ntasks>or-n <ntasks>: How many CPU cores does your job need, in total? -
--time <days-hours:minutes:seconds>or-t <days-hours:minutes:seconds>: How much real-world time (walltime) will your job take to run? The<days>part can be omitted. -
--mem=<megabytes>: How much memory on a node does your job need in megabytes? You can also specify gigabytes using by adding a little “g” afterwards (example:--mem=5g) -
--nodes=<nnodes>or-N <nnodes>: How many separate machines does your job need to run on? Note that if you setntasksto a number greater than what one machine can offer, Slurm will set this value automatically.
Note that just requesting these resources does not make your job run faster, nor does it necessarily mean that you will consume all of these resources. It only means that these are made available to you. Your job may end up using less memory, or less time, or fewer nodes than you have requested, and it will still run.
It’s best if your requests accurately reflect your job’s requirements. We’ll talk more about how to make sure that you’re using resources effectively in a later episode of this lesson.
Submitting Resource Requests
Modify our
hostnamescript so that it runs for a minute, then submit a job for it on the cluster.Solution
[yourUsername@login1 ~]$ cat example-job.sh#!/bin/bash #SBATCH -t 00:01 # timeout in HH:MM echo -n "This script is running on " sleep 20 # time in seconds hostname[yourUsername@login1 ~]$ sbatch example-job.shWhy are the Slurm runtime and
sleeptime not identical?
Resource requests are typically binding. If you exceed them, your job will be killed. Let’s use wall time as an example. We will request 1 minute of wall time, and attempt to run a job for two minutes.
[yourUsername@login1 ~]$ cat example-job.sh
#!/bin/bash
#SBATCH -J long_job
#SBATCH -t 00:01 # timeout in HH:MM
echo "This script is running on ... "
sleep 240 # time in seconds
hostname
Submit the job and wait for it to finish. Once it is has finished, check the log file.
[yourUsername@login1 ~]$ sbatch example-job.sh
[yourUsername@login1 ~]$ squeue -u yourUsername
[yourUsername@login1 ~]$ cat slurm-12.out
This script is running on ...
slurmstepd: error: *** JOB 12 ON node1 CANCELLED AT 2021-02-19T13:55:57
DUE TO TIME LIMIT ***
Our job was killed for exceeding the amount of resources it requested. Although this appears harsh, this is actually a feature. Strict adherence to resource requests allows the scheduler to find the best possible place for your jobs. Even more importantly, it ensures that another user cannot use more resources than they’ve been given. If another user messes up and accidentally attempts to use all of the cores or memory on a node, Slurm will either restrain their job to the requested resources or kill the job outright. Other jobs on the node will be unaffected. This means that one user cannot mess up the experience of others, the only jobs affected by a mistake in scheduling will be their own.
Cancelling a Job
Sometimes we’ll make a mistake and need to cancel a job. This can be done with
the scancel command. Let’s submit a job and then cancel it using
its job number (remember to change the walltime so that it runs long enough for
you to cancel it before it is killed!).
[yourUsername@login1 ~]$ sbatch example-job.sh
[yourUsername@login1 ~]$ squeue -u yourUsername
Submitted batch job 13
JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON)
13 cpubase_b long_job user01 R 0:02 1 node1
Now cancel the job with its job number (printed in your terminal). A clean return of your command prompt indicates that the request to cancel the job was successful.
[yourUsername@login1 ~]$ scancel 38759
# It might take a minute for the job to disappear from the queue...
[yourUsername@login1 ~]$ squeue -u yourUsername
JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON)
Cancelling multiple jobs
We can also cancel all of our jobs at once using the
-uoption. This will delete all jobs for a specific user (in this case, yourself). Note that you can only delete your own jobs.Try submitting multiple jobs and then cancelling them all.
Solution
First, submit a trio of jobs:
[yourUsername@login1 ~]$ sbatch example-job.sh [yourUsername@login1 ~]$ sbatch example-job.sh [yourUsername@login1 ~]$ sbatch example-job.shThen, cancel them all:
[yourUsername@login1 ~]$ scancel -u yourUsername
Other Types of Jobs
Up to this point, we’ve focused on running jobs in batch mode. Slurm also provides the ability to start an interactive session.
There are very frequently tasks that need to be done interactively. Creating an
entire job script might be overkill, but the amount of resources required is
too much for a login node to handle. A good example of this might be building a
genome index for alignment with a tool like HISAT2. Fortunately, we
can run these types of tasks as a one-off with srun.
srun runs a single command on the cluster and then
exits. Let’s demonstrate this by running the hostname command with
srun. (We can cancel an srun
job with Ctrl-c.)
[yourUsername@login1 ~]$ srun hostname
node1
srun accepts all of the same options as
sbatch. However, instead of specifying these in a script,
these options are specified on the command-line when starting a job. To submit
a job that uses 2 CPUs for instance, we could use the following command:
[yourUsername@login1 ~]$ srun -n 2 echo "This job will use 2 CPUs."
This job will use 2 CPUs.
This job will use 2 CPUs.
Typically, the resulting shell environment will be the same as that for
sbatch.
Interactive jobs
Sometimes, you will need a lot of resources for interactive use. Perhaps it’s
our first time running an analysis or we are attempting to debug something that
went wrong with a previous job. Fortunately, Slurm makes it
easy to start an interactive job with srun:
[yourUsername@login1 ~]$ srun --pty bash
You should be presented with a bash prompt. Note that the prompt will likely
change to reflect your new location, in this case the compute node we are
logged on. You can also verify this with hostname.
Creating remote graphics
To see graphical output inside your jobs, you need to use X11 forwarding. To connect with this feature enabled, use the
-Yoption when you login with thesshcommand, e.g.,ssh -Y yourUsername@cluster.hpc-carpentry.org.To demonstrate what happens when you create a graphics window on the remote node, use the
xeyescommand. A relatively adorable pair of eyes should pop up (pressCtrl-Cto stop). If you are using a Mac, you must have installed XQuartz (and restarted your computer) for this to work.If your cluster has the slurm-spank-x11 plugin installed, you can ensure X11 forwarding within interactive jobs by using the
--x11option forsrunwith the commandsrun --x11 --pty bash.
When you are done with the interactive job, type exit to quit your session.
Key Points
The scheduler handles how compute resources are shared between users.
A job is just a shell script.
Request slightly more resources than you will need.
Environment Variables
Overview
Teaching: 10 min
Exercises: 5 minQuestions
How are variables set and accessed in the Unix shell?
How can I use variables to change how a program runs?
Objectives
Understand how variables are implemented in the shell
Read the value of an existing variable
Create new variables and change their values
Change the behaviour of a program using an environment variable
Explain how the shell uses the
PATHvariable to search for executables
Episode provenance
This episode has been remixed from the Shell Extras episode on Shell Variables and the HPC Shell episode on scripts
The shell is just a program, and like other programs, it has variables. Those variables control its execution, so by changing their values you can change how the shell behaves (and with a little more effort how other programs behave).
Variables are a great way of saving information under a name you can access later. In programming languages like Python and R, variables can store pretty much anything you can think of. In the shell, they usually just store text. The best way to understand how they work is to see them in action.
Let’s start by running the command set and looking at some of the variables
in a typical shell session:
$ set
COMPUTERNAME=TURING
HOME=/home/vlad
HOSTNAME=TURING
HOSTTYPE=i686
NUMBER_OF_PROCESSORS=4
PATH=/Users/vlad/bin:/usr/local/git/bin:/usr/bin:/bin:/usr/sbin:/sbin:/usr/local/bin
PWD=/home/vlad
UID=1000
USERNAME=vlad
...
As you can see, there are quite a few — in fact,
four or five times more than what’s shown here.
And yes, using set to show things might seem a little strange,
even for Unix, but if you don’t give it any arguments,
it might as well show you things you could set.
Every variable has a name.
All shell variables’ values are strings,
even those (like UID) that look like numbers.
It’s up to programs to convert these strings to other types when necessary.
For example, if a program wanted to find out how many processors the computer
had, it would convert the value of the NUMBER_OF_PROCESSORS variable from a
string to an integer.
Showing the Value of a Variable
Let’s show the value of the variable HOME:
$ echo HOME
HOME
That just prints “HOME”, which isn’t what we wanted (though it is what we actually asked for). Let’s try this instead:
$ echo $HOME
/home/vlad
The dollar sign tells the shell that we want the value of the variable
rather than its name.
This works just like wildcards:
the shell does the replacement before running the program we’ve asked for.
Thanks to this expansion, what we actually run is echo /home/vlad,
which displays the right thing.
Creating and Changing Variables
Creating a variable is easy — we just assign a value to a name using “=”
(we just have to remember that the syntax requires that there are no spaces
around the =!):
$ SECRET_IDENTITY=Dracula
$ echo $SECRET_IDENTITY
Dracula
To change the value, just assign a new one:
$ SECRET_IDENTITY=Camilla
$ echo $SECRET_IDENTITY
Camilla
Environment variables
When we ran the set command we saw there were a lot of variables whose names
were in upper case. That’s because, by convention, variables that are also
available to use by other programs are given upper-case names. Such variables
are called environment variables as they are shell variables that are defined
for the current shell and are inherited by any child shells or processes.
To create an environment variable you need to export a shell variable. For
example, to make our SECRET_IDENTITY available to other programs that we call
from our shell we can do:
$ SECRET_IDENTITY=Camilla
$ export SECRET_IDENTITY
You can also create and export the variable in a single step:
$ export SECRET_IDENTITY=Camilla
Using environment variables to change program behaviour
Set a shell variable
TIME_STYLEto have a value ofisoand check this value using theechocommand.Now, run the command
lswith the option-l(which gives a long format).
exportthe variable and rerun thels -lcommand. Do you notice any difference?Solution
The
TIME_STYLEvariable is not seen bylsuntil is exported, at which point it is used bylsto decide what date format to use when presenting the timestamp of files.
You can see the complete set of environment variables in your current shell
session with the command env (which returns a subset of what the command
set gave us). The complete set of environment variables is called
your runtime environment and can affect the behaviour of the programs you
run.
Job environment variables
When Slurm runs a job, it sets a number of environment variables for the job. One of these will let us check what directory our job script was submitted from. The
SLURM_SUBMIT_DIRvariable is set to the directory from which our job was submitted. Using theSLURM_SUBMIT_DIRvariable, modify your job so that it prints out the location from which the job was submitted.Solution
[yourUsername@login1 ~]$ nano example-job.sh [yourUsername@login1 ~]$ cat example-job.sh#!/bin/bash #SBATCH -t 00:00:30 echo -n "This script is running on " hostname echo "This job was launched in the following directory:" echo ${SLURM_SUBMIT_DIR}
To remove a variable or environment variable you can use the unset command,
for example:
$ unset SECRET_IDENTITY
The PATH Environment Variable
Similarly, some environment variables (like PATH) store lists of values.
In this case, the convention is to use a colon ‘:’ as a separator.
If a program wants the individual elements of such a list,
it’s the program’s responsibility to split the variable’s string value into
pieces.
Let’s have a closer look at that PATH variable.
Its value defines the shell’s search path for executables,
i.e., the list of directories that the shell looks in for runnable programs
when you type in a program name without specifying what directory it is in.
For example, when we type a command like analyze,
the shell needs to decide whether to run ./analyze or /bin/analyze.
The rule it uses is simple:
the shell checks each directory in the PATH variable in turn,
looking for a program with the requested name in that directory.
As soon as it finds a match, it stops searching and runs the program.
To show how this works,
here are the components of PATH listed one per line:
/Users/vlad/bin
/usr/local/git/bin
/usr/bin
/bin
/usr/sbin
/sbin
/usr/local/bin
On our computer,
there are actually three programs called analyze
in three different directories:
/bin/analyze,
/usr/local/bin/analyze,
and /users/vlad/analyze.
Since the shell searches the directories in the order they’re listed in PATH,
it finds /bin/analyze first and runs that.
Notice that it will never find the program /users/vlad/analyze
unless we type in the full path to the program,
since the directory /users/vlad isn’t in PATH.
This means that I can have executables in lots of different places as long as
I remember that I need to to update my PATH so that my shell can find them.
What if I want to run two different versions of the same program?
Since they share the same name, if I add them both to my PATH the first one
found will always win.
In the next episode we’ll learn how to use helper tools to help us manage our
runtime environment to make that possible without us needing to do a lot of
bookkeeping on what the value of PATH (and other important environment
variables) is or should be.
Key Points
Shell variables are by default treated as strings
Variables are assigned using “
=” and recalled using the variable’s name prefixed by “$”Use “
export” to make an variable available to other programsThe
PATHvariable defines the shell’s search path
Accessing software via Modules
Overview
Teaching: 30 min
Exercises: 15 minQuestions
How do we load and unload software packages?
Objectives
Load and use a software package.
Explain how the shell environment changes when the module mechanism loads or unloads packages.
On a high-performance computing system, it is seldom the case that the software we want to use is available when we log in. It is installed, but we will need to “load” it before it can run.
Before we start using individual software packages, however, we should understand the reasoning behind this approach. The three biggest factors are:
- software incompatibilities
- versioning
- dependencies
Software incompatibility is a major headache for programmers. Sometimes the
presence (or absence) of a software package will break others that depend on
it. Two well known examples are Python and C compiler versions.
Python 3 famously provides a python command that conflicts with that provided
by Python 2. Software compiled against a newer version of the C libraries and
then run on a machine that has older C libraries installed will result in a
nasty 'GLIBCXX_3.4.20' not found error.
Software versioning is another common issue. A team might depend on a certain package version for their research project - if the software version was to change (for instance, if a package was updated), it might affect their results. Having access to multiple software versions allows a set of researchers to prevent software versioning issues from affecting their results.
Dependencies are where a particular software package (or even a particular version) depends on having access to another software package (or even a particular version of another software package). For example, the VASP materials science software may depend on having a particular version of the FFTW (Fastest Fourier Transform in the West) software library available for it to work.
Environment Modules
Environment modules are the solution to these problems. A module is a self-contained description of a software package – it contains the settings required to run a software package and, usually, encodes required dependencies on other software packages.
There are a number of different environment module implementations commonly
used on HPC systems: the two most common are TCL modules and Lmod. Both of
these use similar syntax and the concepts are the same so learning to use one
will allow you to use whichever is installed on the system you are using. In
both implementations the module command is used to interact with environment
modules. An additional subcommand is usually added to the command to specify
what you want to do. For a list of subcommands you can use module -h or
module help. As for all commands, you can access the full help on the man
pages with man module.
On login you may start out with a default set of modules loaded or you may start out with an empty environment; this depends on the setup of the system you are using.
Listing Available Modules
To see available software modules, use module avail:
[yourUsername@login1 ~]$ module avail
~~~ /cvmfs/pilot.eessi-hpc.org/2020.12/software/x86_64/amd/zen2/modules/all ~~~
Bazel/3.6.0-GCCcore-x.y.z NSS/3.51-GCCcore-x.y.z
Bison/3.5.3-GCCcore-x.y.z Ninja/1.10.0-GCCcore-x.y.z
Boost/1.72.0-gompi-2020a OSU-Micro-Benchmarks/5.6.3-gompi-2020a
CGAL/4.14.3-gompi-2020a-Python-3.x.y OpenBLAS/0.3.9-GCC-x.y.z
CMake/3.16.4-GCCcore-x.y.z OpenFOAM/v2006-foss-2020a
[removed most of the output here for clarity]
Where:
L: Module is loaded
Aliases: Aliases exist: foo/1.2.3 (1.2) means that "module load foo/1.2"
will load foo/1.2.3
D: Default Module
Use "module spider" to find all possible modules and extensions.
Use "module keyword key1 key2 ..." to search for all possible modules matching
any of the "keys".
Listing Currently Loaded Modules
You can use the module list command to see which modules you currently have
loaded in your environment. If you have no modules loaded, you will see a
message telling you so
[yourUsername@login1 ~]$ module list
No Modulefiles Currently Loaded.
Loading and Unloading Software
To load a software module, use module load. In this example we will use
Python 3.
Initially, Python 3 is not loaded. We can test this by using the which
command. which looks for programs the same way that Bash does, so we can use
it to tell us where a particular piece of software is stored.
[yourUsername@login1 ~]$ which python3
If the python3 command was unavailable, we would see output like
/usr/bin/which: no python3 in (/cvmfs/pilot.eessi-hpc.org/2020.12/compat/linux/x86_64/usr/bin:/opt/software/slurm/bin:/usr/local/bin:/usr/bin:/usr/local/sbin:/usr/sbin:/opt/puppetlabs/bin:/home/yourUsername/.local/bin:/home/yourUsername/bin)
Note that this wall of text is really a list, with values separated
by the : character. The output is telling us that the which command
searched the following directories for python3, without success:
/cvmfs/pilot.eessi-hpc.org/2020.12/compat/linux/x86_64/usr/bin
/opt/software/slurm/bin
/usr/local/bin
/usr/bin
/usr/local/sbin
/usr/sbin
/opt/puppetlabs/bin
/home/yourUsername/.local/bin
/home/yourUsername/bin
However, in our case we do have an existing python3 available so we see
/cvmfs/pilot.eessi-hpc.org/2020.12/compat/linux/x86_64/usr/bin/python3
We need a different Python than the system provided one though, so let us load a module to access it.
We can load the python3 command with module load:
[yourUsername@login1 ~]$ module load Python
[yourUsername@login1 ~]$ which python3
/cvmfs/pilot.eessi-hpc.org/2020.12/software/x86_64/amd/zen2/software/Python/3.x.y-GCCcore-x.y.z/bin/python3
So, what just happened?
To understand the output, first we need to understand the nature of the $PATH
environment variable. $PATH is a special environment variable that controls
where a UNIX system looks for software. Specifically $PATH is a list of
directories (separated by :) that the OS searches through for a command
before giving up and telling us it can’t find it. As with all environment
variables we can print it out using echo.
[yourUsername@login1 ~]$ echo $PATH
/cvmfs/pilot.eessi-hpc.org/2020.12/software/x86_64/amd/zen2/software/Python/3.x.y-GCCcore-x.y.z/bin:/cvmfs/pilot.eessi-hpc.org/2020.12/software/x86_64/amd/zen2/software/SQLite/3.31.1-GCCcore-x.y.z/bin:/cvmfs/pilot.eessi-hpc.org/2020.12/software/x86_64/amd/zen2/software/Tcl/8.6.10-GCCcore-x.y.z/bin:/cvmfs/pilot.eessi-hpc.org/2020.12/software/x86_64/amd/zen2/software/GCCcore/x.y.z/bin:/cvmfs/pilot.eessi-hpc.org/2020.12/compat/linux/x86_64/usr/bin:/opt/software/slurm/bin:/usr/local/bin:/usr/bin:/usr/local/sbin:/usr/sbin:/opt/puppetlabs/bin:/home/user01/.local/bin:/home/user01/bin
You’ll notice a similarity to the output of the which command. In this case,
there’s only one difference: the different directory at the beginning. When we
ran the module load command, it added a directory to the beginning of our
$PATH. Let’s examine what’s there:
[yourUsername@login1 ~]$ ls /cvmfs/pilot.eessi-hpc.org/2020.12/software/x86_64/amd/zen2/software/Python/3.x.y-GCCcore-x.y.z/bin
2to3 nosetests-3.8 python rst2s5.py
2to3-3.8 pasteurize python3 rst2xetex.py
chardetect pbr python3.8 rst2xml.py
cygdb pip python3.8-config rstpep2html.py
cython pip3 python3-config runxlrd.py
cythonize pip3.8 rst2html4.py sphinx-apidoc
easy_install pybabel rst2html5.py sphinx-autogen
easy_install-3.8 __pycache__ rst2html.py sphinx-build
futurize pydoc3 rst2latex.py sphinx-quickstart
idle3 pydoc3.8 rst2man.py tabulate
idle3.8 pygmentize rst2odt_prepstyles.py virtualenv
netaddr pytest rst2odt.py wheel
nosetests py.test rst2pseudoxml.py
Taking this to its conclusion, module load will add software to your $PATH.
It “loads” software. A special note on this - depending on which version of the
module program that is installed at your site, module load will also load
required software dependencies.
To demonstrate, let’s use module list. module list shows all loaded
software modules.
[yourUsername@login1 ~]$ module list
Currently Loaded Modules:
1) GCCcore/x.y.z 4) GMP/6.2.0-GCCcore-x.y.z
2) Tcl/8.6.10-GCCcore-x.y.z 5) libffi/3.3-GCCcore-x.y.z
3) SQLite/3.31.1-GCCcore-x.y.z 6) Python/3.x.y-GCCcore-x.y.z
[yourUsername@login1 ~]$ module load GROMACS
[yourUsername@login1 ~]$ module list
Currently Loaded Modules:
1) GCCcore/x.y.z 14) libfabric/1.11.0-GCCcore-x.y.z
2) Tcl/8.6.10-GCCcore-x.y.z 15) PMIx/3.1.5-GCCcore-x.y.z
3) SQLite/3.31.1-GCCcore-x.y.z 16) OpenMPI/4.0.3-GCC-x.y.z
4) GMP/6.2.0-GCCcore-x.y.z 17) OpenBLAS/0.3.9-GCC-x.y.z
5) libffi/3.3-GCCcore-x.y.z 18) gompi/2020a
6) Python/3.x.y-GCCcore-x.y.z 19) FFTW/3.3.8-gompi-2020a
7) GCC/x.y.z 20) ScaLAPACK/2.1.0-gompi-2020a
8) numactl/2.0.13-GCCcore-x.y.z 21) foss/2020a
9) libxml2/2.9.10-GCCcore-x.y.z 22) pybind11/2.4.3-GCCcore-x.y.z-Pytho...
10) libpciaccess/0.16-GCCcore-x.y.z 23) SciPy-bundle/2020.03-foss-2020a-Py...
11) hwloc/2.2.0-GCCcore-x.y.z 24) networkx/2.4-foss-2020a-Python-3.8...
12) libevent/2.1.11-GCCcore-x.y.z 25) GROMACS/2020.1-foss-2020a-Python-3...
13) UCX/1.8.0-GCCcore-x.y.z
So in this case, loading the GROMACS module (a bioinformatics software
package), also loaded GMP/6.2.0-GCCcore-x.y.z and
SciPy-bundle/2020.03-foss-2020a-Python-3.x.y as well. Let’s try unloading the
GROMACS package.
[yourUsername@login1 ~]$ module unload GROMACS
[yourUsername@login1 ~]$ module list
Currently Loaded Modules:
1) GCCcore/x.y.z 13) UCX/1.8.0-GCCcore-x.y.z
2) Tcl/8.6.10-GCCcore-x.y.z 14) libfabric/1.11.0-GCCcore-x.y.z
3) SQLite/3.31.1-GCCcore-x.y.z 15) PMIx/3.1.5-GCCcore-x.y.z
4) GMP/6.2.0-GCCcore-x.y.z 16) OpenMPI/4.0.3-GCC-x.y.z
5) libffi/3.3-GCCcore-x.y.z 17) OpenBLAS/0.3.9-GCC-x.y.z
6) Python/3.x.y-GCCcore-x.y.z 18) gompi/2020a
7) GCC/x.y.z 19) FFTW/3.3.8-gompi-2020a
8) numactl/2.0.13-GCCcore-x.y.z 20) ScaLAPACK/2.1.0-gompi-2020a
9) libxml2/2.9.10-GCCcore-x.y.z 21) foss/2020a
10) libpciaccess/0.16-GCCcore-x.y.z 22) pybind11/2.4.3-GCCcore-x.y.z-Pytho...
11) hwloc/2.2.0-GCCcore-x.y.z 23) SciPy-bundle/2020.03-foss-2020a-Py...
12) libevent/2.1.11-GCCcore-x.y.z 24) networkx/2.4-foss-2020a-Python-3.x.y
So using module unload “un-loads” a module, and depending on how a site is
configured it may also unload all of the dependencies (in our case it does
not). If we wanted to unload everything at once, we could run module purge
(unloads everything).
[yourUsername@login1 ~]$ module purge
[yourUsername@login1 ~]$ module list
No modules loaded
Note that module purge is informative. It will also let us know if a default
set of “sticky” packages cannot be unloaded (and how to actually unload these
if we truly so desired).
Note that this module loading process happens principally through
the manipulation of environment variables like $PATH. There
is usually little or no data transfer involved.
The module loading process manipulates other special environment variables as well, including variables that influence where the system looks for software libraries, and sometimes variables which tell commercial software packages where to find license servers.
The module command also restores these shell environment variables to their previous state when a module is unloaded.
Software Versioning
So far, we’ve learned how to load and unload software packages. This is very useful. However, we have not yet addressed the issue of software versioning. At some point or other, you will run into issues where only one particular version of some software will be suitable. Perhaps a key bugfix only happened in a certain version, or version X broke compatibility with a file format you use. In either of these example cases, it helps to be very specific about what software is loaded.
Let’s examine the output of module avail more closely.
[yourUsername@login1 ~]$ module avail
~~~ /cvmfs/pilot.eessi-hpc.org/2020.12/software/x86_64/amd/zen2/modules/all ~~~
Bazel/3.6.0-GCCcore-x.y.z NSS/3.51-GCCcore-x.y.z
Bison/3.5.3-GCCcore-x.y.z Ninja/1.10.0-GCCcore-x.y.z
Boost/1.72.0-gompi-2020a OSU-Micro-Benchmarks/5.6.3-gompi-2020a
CGAL/4.14.3-gompi-2020a-Python-3.x.y OpenBLAS/0.3.9-GCC-x.y.z
CMake/3.16.4-GCCcore-x.y.z OpenFOAM/v2006-foss-2020a
[removed most of the output here for clarity]
Where:
L: Module is loaded
Aliases: Aliases exist: foo/1.2.3 (1.2) means that "module load foo/1.2"
will load foo/1.2.3
D: Default Module
Use "module spider" to find all possible modules and extensions.
Use "module keyword key1 key2 ..." to search for all possible modules matching
any of the "keys".
Using Software Modules in Scripts
Create a job that is able to run
python3 --version. Remember, no software is loaded by default! Running a job is just like logging on to the system (you should not assume a module loaded on the login node is loaded on a compute node).Solution
[yourUsername@login1 ~]$ nano python-module.sh [yourUsername@login1 ~]$ cat python-module.sh#!/bin/bash #SBATCH #SBATCH -t 00:00:30 module load Python python3 --version[yourUsername@login1 ~]$ sbatch python-module.sh
Key Points
Load software with
module load softwareName.Unload software with
module unloadThe module system handles software versioning and package conflicts for you automatically.
Transferring files with remote computers
Overview
Teaching: 15 min
Exercises: 15 minQuestions
How do I transfer files to (and from) the cluster?
Objectives
Transfer files to and from a computing cluster.
Performing work on a remote computer is not very useful if we cannot get files to or from the cluster. There are several options for transferring data between computing resources using CLI and GUI utilities, a few of which we will cover.
Download Lesson Files From the Internet
One of the most straightforward ways to download files is to use either curl
or wget. One of these is usually installed in most Linux shells, on Mac OS
terminal and in GitBash. Any file that can be downloaded in your web browser
through a direct link can be downloaded using curl or wget. This is a
quick way to download datasets or source code. The syntax for these commands is
wget [-O new_name] https://some/link/to/a/filecurl [-o new_name] https://some/link/to/a/file
Try it out by downloading some material we’ll use later on, from a terminal on your local machine, using the URL of the current codebase:
https://github.com/hpc-carpentry/amdahl/tarball/main
Download the “Tarball”
The word “tarball” in the above URL refers to a compressed archive format commonly used on Linux, which is the operating system the majority of HPC cluster machines run. A tarball is a lot like a
.zipfile. The actual file extension is.tar.gz, which reflects the two-stage process used to create the file: the files or folders are merged into a single file usingtar, which is then compressed usinggzip, so the file extension is “tar-dot-g-z.” That’s a mouthful, so people often say “the xyz tarball” instead.You may also see the extension
.tgz, which is just an abbreviation of.tar.gz.By default,
curlandwgetdownload files to the same name as the URL: in this case,main. Use one of the above commands to save the tarball asamdahl.tar.gz.
wgetandcurlCommands[you@laptop:~]$ wget -O amdahl.tar.gz https://github.com/hpc-carpentry/amdahl/tarball/main # or [you@laptop:~]$ curl -o amdahl.tar.gz -L https://github.com/hpc-carpentry/amdahl/tarball/mainThe
-Loption tocurltells it to follow URL redirects (whichwgetdoes by default).
After downloading the file, use ls to see it in your working directory:
[you@laptop:~]$ ls
Archiving Files
One of the biggest challenges we often face when transferring data between remote HPC systems is that of large numbers of files. There is an overhead to transferring each individual file and when we are transferring large numbers of files these overheads combine to slow down our transfers to a large degree.
The solution to this problem is to archive multiple files into smaller
numbers of larger files before we transfer the data to improve our transfer
efficiency.
Sometimes we will combine archiving with compression to reduce the amount of
data we have to transfer and so speed up the transfer.
The most common archiving command you will use on a (Linux) HPC cluster is
tar.
tar can be used to combine files and folders into a single archive file and,
optionally, compress the result.
Let’s look at the file we downloaded from the lesson site, amdahl.tar.gz.
The .gz part stands for gzip, which is a compression library.
It’s common (but not necessary!) that this kind of file can be interpreted by
reading its name: it appears somebody took files and folders relating to
something called “amdahl,” wrapped them all up into a single file with tar,
then compressed that archive with gzip to save space.
Let’s see if that is the case, without unpacking the file.
tar prints the “table of contents” with the -t flag, for the file
specified with the -f flag followed by the filename.
Note that you can concatenate the two flags: writing -t -f is interchangeable
with writing -tf together.
However, the argument following -f must be a filename, so writing -ft will
not work.
[you@laptop:~]$ tar -tf amdahl.tar.gz
hpc-carpentry-amdahl-46c9b4b/
hpc-carpentry-amdahl-46c9b4b/.github/
hpc-carpentry-amdahl-46c9b4b/.github/workflows/
hpc-carpentry-amdahl-46c9b4b/.github/workflows/python-publish.yml
hpc-carpentry-amdahl-46c9b4b/.gitignore
hpc-carpentry-amdahl-46c9b4b/LICENSE
hpc-carpentry-amdahl-46c9b4b/README.md
hpc-carpentry-amdahl-46c9b4b/amdahl/
hpc-carpentry-amdahl-46c9b4b/amdahl/__init__.py
hpc-carpentry-amdahl-46c9b4b/amdahl/__main__.py
hpc-carpentry-amdahl-46c9b4b/amdahl/amdahl.py
hpc-carpentry-amdahl-46c9b4b/requirements.txt
hpc-carpentry-amdahl-46c9b4b/setup.py
This example output shows a folder which contains a few files, where 46c9b4b
is an 8-character git commit hash that will change when the source
material is updated.
Now let’s unpack the archive. We’ll run tar with a few common flags:
-xto extract the archive-vfor verbose output-zfor gzip compression-f «tarball»for the file to be unpacked
Extract the Archive
Using the flags above, unpack the source code tarball into a new directory named “amdahl” using
tar.[you@laptop:~]$ tar -xvzf amdahl.tar.gzhpc-carpentry-amdahl-46c9b4b/ hpc-carpentry-amdahl-46c9b4b/.github/ hpc-carpentry-amdahl-46c9b4b/.github/workflows/ hpc-carpentry-amdahl-46c9b4b/.github/workflows/python-publish.yml hpc-carpentry-amdahl-46c9b4b/.gitignore hpc-carpentry-amdahl-46c9b4b/LICENSE hpc-carpentry-amdahl-46c9b4b/README.md hpc-carpentry-amdahl-46c9b4b/amdahl/ hpc-carpentry-amdahl-46c9b4b/amdahl/__init__.py hpc-carpentry-amdahl-46c9b4b/amdahl/__main__.py hpc-carpentry-amdahl-46c9b4b/amdahl/amdahl.py hpc-carpentry-amdahl-46c9b4b/requirements.txt hpc-carpentry-amdahl-46c9b4b/setup.pyNote that we did not need to type out
-x -v -z -f, thanks to flag concatenation, though the command works identically either way – so long as the concatenated list ends withf, because the next string must specify the name of the file to extract.
The folder has an unfortunate name, so let’s change that to something more convenient.
[you@laptop:~]$ mv hpc-carpentry-amdahl-46c9b4b amdahl
Check the size of the extracted directory and compare to the compressed
file size, using du for “disk usage”.
[you@laptop:~]$ du -sh amdahl.tar.gz
8.0K amdahl.tar.gz
[you@laptop:~]$ du -sh amdahl
48K amdahl
Text files (including Python source code) compress nicely: the “tarball” is one-sixth the total size of the raw data!
If you want to reverse the process – compressing raw data instead of
extracting it – set a c flag instead of x, set the archive filename,
then provide a directory to compress:
[you@laptop:~]$ tar -cvzf compressed_code.tar.gz amdahl
amdahl/
amdahl/.github/
amdahl/.github/workflows/
amdahl/.github/workflows/python-publish.yml
amdahl/.gitignore
amdahl/LICENSE
amdahl/README.md
amdahl/amdahl/
amdahl/amdahl/__init__.py
amdahl/amdahl/__main__.py
amdahl/amdahl/amdahl.py
amdahl/requirements.txt
amdahl/setup.py
If you give amdahl.tar.gz as the filename in the above command, tar will
update the existing tarball with any changes you made to the files.
That would mean adding the new amdahl folder to the existing folder
(hpc-carpentry-amdahl-46c9b4b) inside the tarball, doubling the size of the
archive!
Working with Windows
When you transfer text files from a Windows system to a Unix system (Mac, Linux, BSD, Solaris, etc.) this can cause problems. Windows encodes its files slightly different than Unix, and adds an extra character to every line.
On a Unix system, every line in a file ends with a
\n(newline). On Windows, every line in a file ends with a\r\n(carriage return + newline). This causes problems sometimes.Though most modern programming languages and software handles this correctly, in some rare instances, you may run into an issue. The solution is to convert a file from Windows to Unix encoding with the
dos2unixcommand.You can identify if a file has Windows line endings with
cat -A filename. A file with Windows line endings will have^M$at the end of every line. A file with Unix line endings will have$at the end of a line.To convert the file, just run
dos2unix filename. (Conversely, to convert back to Windows format, you can rununix2dos filename.)
Transferring Single Files and Folders With scp
To copy a single file to or from the cluster, we can use scp (“secure copy”).
The syntax can be a little complex for new users, but we’ll break it down.
The scp command is a relative of the ssh command we used to
access the system, and can use the same public-key authentication
mechanism.
To upload to another computer, the template command is
[you@laptop:~]$ scp local_file yourUsername@cluster.hpc-carpentry.org:remote_destination
in which @ and : are field separators and remote_destination is a path
relative to your remote home directory, or a new filename if you wish to change
it, or both a relative path and a new filename.
If you don’t have a specific folder in mind you can omit the
remote_destination and the file will be copied to your home directory on the
remote computer (with its original name).
If you include a remote_destination, note that scp interprets this the same
way cp does when making local copies:
if it exists and is a folder, the file is copied inside the folder; if it
exists and is a file, the file is overwritten with the contents of
local_file; if it does not exist, it is assumed to be a destination filename
for local_file.
Upload the lesson material to your remote home directory like so:
[you@laptop:~]$ scp amdahl.tar.gz yourUsername@cluster.hpc-carpentry.org:
Why Not Download on HPC Carpentry’s Cloud Cluster Directly?
Most computer clusters are protected from the open internet by a firewall. For enhanced security, some are configured to allow traffic inbound, but not outbound. This means that an authenticated user can send a file to a cluster machine, but a cluster machine cannot retrieve files from a user’s machine or the open Internet.
Try downloading the file directly. Note that it may well fail, and that’s OK!
Commands
[you@laptop:~]$ ssh yourUsername@cluster.hpc-carpentry.org [yourUsername@login1 ~]$ wget -O amdahl.tar.gz https://github.com/hpc-carpentry/amdahl/tarball/main # or [yourUsername@login1 ~]$ curl -o amdahl.tar.gz https://github.com/hpc-carpentry/amdahl/tarball/mainDid it work? If not, what does the terminal output tell you about what happened?
Transferring a Directory
To transfer an entire directory, we add the -r flag for “recursive”:
copy the item specified, and every item below it, and every item below those…
until it reaches the bottom of the directory tree rooted at the folder name you
provided.
[you@laptop:~]$ scp -r amdahl yourUsername@cluster.hpc-carpentry.org:
Caution
For a large directory – either in size or number of files – copying with
-rcan take a long time to complete.
When using scp, you may have noticed that a : always follows the remote
computer name.
A string after the : specifies the remote directory you wish to transfer
the file or folder to, including a new name if you wish to rename the remote
material.
If you leave this field blank, scp defaults to your home directory and the
name of the local material to be transferred.
On Linux computers, / is the separator in file or directory paths.
A path starting with a / is called absolute, since there can be nothing
above the root /.
A path that does not start with / is called relative, since it is not
anchored to the root.
If you want to upload a file to a location inside your home directory –
which is often the case – then you don’t need a leading /. After the :,
you can type the destination path relative to your home directory.
If your home directory is the destination, you can leave the destination
field blank, or type ~ – the shorthand for your home directory – for
completeness.
With scp, a trailing slash on the target directory is optional, and has no effect.
A trailing slash on a source directory is important for other commands, like rsync.
A Note on
rsyncAs you gain experience with transferring files, you may find the
scpcommand limiting. The rsync utility provides advanced features for file transfer and is typically faster compared to bothscpandsftp(see below). It is especially useful for transferring large and/or many files and for synchronizing folder contents between computers.The syntax is similar to
scp. To transfer to another computer with commonly used options:[you@laptop:~]$ rsync -avP amdahl.tar.gz yourUsername@cluster.hpc-carpentry.org:The options are:
-a(archive) to preserve file timestamps, permissions, and folders, among other things; implies recursion-v(verbose) to get verbose output to help monitor the transfer-P(partial/progress) to preserve partially transferred files in case of an interruption and also displays the progress of the transfer.To recursively copy a directory, we can use the same options:
[you@laptop:~]$ rsync -avP amdahl yourUsername@cluster.hpc-carpentry.org:~/As written, this will place the local directory and its contents under your home directory on the remote system. If a trailing slash is added to the source, a new directory corresponding to the transferred directory will not be created, and the contents of the source directory will be copied directly into the destination directory.
To download a file, we simply change the source and destination:
[you@laptop:~]$ rsync -avP yourUsername@cluster.hpc-carpentry.org:amdahl ./
File transfers using both scp and rsync use SSH to encrypt data sent through
the network. So, if you can connect via SSH, you will be able to transfer
files. By default, SSH uses network port 22. If a custom SSH port is in use,
you will have to specify it using the appropriate flag, often -p, -P, or
--port. Check --help or the man page if you’re unsure.
Change the Rsync Port
Say we have to connect
rsyncthrough port 768 instead of 22. How would we modify this command?[you@laptop:~]$ rsync amdahl.tar.gz yourUsername@cluster.hpc-carpentry.org:Hint: check the
manpage or “help” forrsync.Solution
[you@laptop:~]$ man rsync [you@laptop:~]$ rsync --help | grep port --port=PORT specify double-colon alternate port number See http://rsync.samba.org/ for updates, bug reports, and answers [you@laptop:~]$ rsync --port=768 amdahl.tar.gz yourUsername@cluster.hpc-carpentry.org:(Note that this command will fail, as the correct port in this case is the default: 22.)
Transferring Files Interactively with FileZilla
FileZilla is a cross-platform client for downloading and uploading files to and
from a remote computer. It is absolutely fool-proof and always works quite
well. It uses the sftp protocol. You can read more about using the sftp
protocol in the command line in the
lesson discussion.
Download and install the FileZilla client from https://filezilla-project.org. After installing and opening the program, you should end up with a window with a file browser of your local system on the left hand side of the screen. When you connect to the cluster, your cluster files will appear on the right hand side.
To connect to the cluster, we’ll just need to enter our credentials at the top of the screen:
- Host:
sftp://cluster.hpc-carpentry.org - User: Your cluster username
- Password: Your cluster password
- Port: (leave blank to use the default port)
Hit “Quickconnect” to connect. You should see your remote files appear on the right hand side of the screen. You can drag-and-drop files between the left (local) and right (remote) sides of the screen to transfer files.
Finally, if you need to move large files (typically larger than a gigabyte)
from one remote computer to another remote computer, SSH in to the computer
hosting the files and use scp or rsync to transfer over to the other. This
will be more efficient than using FileZilla (or related applications) that
would copy from the source to your local machine, then to the destination
machine.
Key Points
wgetandcurl -Odownload a file from the internet.
scpandrsynctransfer files to and from your computer.You can use an SFTP client like FileZilla to transfer files through a GUI.
Running a parallel job
Overview
Teaching: 30 min
Exercises: 60 minQuestions
How do we execute a task in parallel?
What benefits arise from parallel execution?
What are the limits of gains from execution in parallel?
Objectives
Install a Python package using
pipPrepare a job submission script for the parallel executable.
Launch jobs with parallel execution.
Record and summarize the timing and accuracy of jobs.
Describe the relationship between job parallelism and performance.
We now have the tools we need to run a multi-processor job. This is a very important aspect of HPC systems, as parallelism is one of the primary tools we have to improve the performance of computational tasks.
If you disconnected, log back in to the cluster.
[you@laptop:~]$ ssh yourUsername@cluster.hpc-carpentry.org
Install the Amdahl Program
With the Amdahl source code on the cluster, we can install it, which will
provide access to the amdahl executable.
Move into the extracted directory, then use the Package Installer for Python,
or pip, to install it in your (“user”) home directory:
[yourUsername@login1 ~]$ cd amdahl
[yourUsername@login1 ~]$ python3 -m pip install --user .
Amdahl is Python Code
The Amdahl program is written in Python, and installing or using it requires locating the
python3executable on the login node. If it can’t be found, try listing available modules usingmodule avail, load the appropriate one, and try the command again.
MPI for Python
The Amdahl code has one dependency: mpi4py.
If it hasn’t already been installed on the cluster, pip will attempt to
collect mpi4py from the Internet and install it for you.
If this fails due to a one-way firewall, you must retrieve mpi4py on your
local machine and upload it, just as we did for Amdahl.
Retrieve and Upload
mpi4pyIf installing Amdahl failed because mpi4py could not be installed, retrieve the tarball from https://github.com/mpi4py/mpi4py/tarball/master then
rsyncit to the cluster, extract, and install:[you@laptop:~]$ wget -O mpi4py.tar.gz https://github.com/mpi4py/mpi4py/releases/download/3.1.4/mpi4py-3.1.4.tar.gz [you@laptop:~]$ scp mpi4py.tar.gz yourUsername@cluster.hpc-carpentry.org: # or [you@laptop:~]$ rsync -avP mpi4py.tar.gz yourUsername@cluster.hpc-carpentry.org:[you@laptop:~]$ ssh yourUsername@cluster.hpc-carpentry.org [yourUsername@login1 ~]$ tar -xvzf mpi4py.tar.gz # extract the archive [yourUsername@login1 ~]$ mv mpi4py* mpi4py # rename the directory [yourUsername@login1 ~]$ cd mpi4py [yourUsername@login1 ~]$ python3 -m pip install --user . [yourUsername@login1 ~]$ cd ../amdahl [yourUsername@login1 ~]$ python3 -m pip install --user .
If
pipRaises a Warning…
pipmay warn that your user package binaries are not in your PATH.WARNING: The script amdahl is installed in "${HOME}/.local/bin" which is not on PATH. Consider adding this directory to PATH or, if you prefer to suppress this warning, use --no-warn-script-location.To check whether this warning is a problem, use
whichto search for theamdahlprogram:[yourUsername@login1 ~]$ which amdahlIf the command returns no output, displaying a new prompt, it means the file
amdahlhas not been found. You must update the environment variable namedPATHto include the missing folder. Edit your shell configuration file as follows, then log off the cluster and back on again so it takes effect.[yourUsername@login1 ~]$ nano ~/.bashrc [yourUsername@login1 ~]$ tail ~/.bashrcexport PATH=${PATH}:${HOME}/.local/binAfter logging back in to cluster.hpc-carpentry.org,
whichshould be able to findamdahlwithout difficulties. If you had to load a Python module, load it again.
Help!
Many command-line programs include a “help” message. Try it with amdahl:
[yourUsername@login1 ~]$ amdahl --help
usage: amdahl [-h] [-p [PARALLEL_PROPORTION]] [-w [WORK_SECONDS]] [-t] [-e] [-j [JITTER_PROPORTION]]
optional arguments:
-h, --help show this help message and exit
-p [PARALLEL_PROPORTION], --parallel-proportion [PARALLEL_PROPORTION]
Parallel proportion: a float between 0 and 1
-w [WORK_SECONDS], --work-seconds [WORK_SECONDS]
Total seconds of workload: an integer greater than 0
-t, --terse Format output as a machine-readable object for easier analysis
-e, --exact Exactly match requested timing by disabling random jitter
-j [JITTER_PROPORTION], --jitter-proportion [JITTER_PROPORTION]
Random jitter: a float between -1 and +1
This message doesn’t tell us much about what the program does, but it does tell us the important flags we might want to use when launching it.
Running the Job on a Compute Node
Create a submission file, requesting one task on a single node, then launch it.
[yourUsername@login1 ~]$ nano serial-job.sh
[yourUsername@login1 ~]$ cat serial-job.sh
#!/bin/bash
#SBATCH -J solo-job
#SBATCH -p cpubase_bycore_b1
#SBATCH -N 1
#SBATCH -n 1
# Load the computing environment we need
module load Python
# Execute the task
amdahl
[yourUsername@login1 ~]$ sbatch serial-job.sh
As before, use the Slurm status commands to check whether your job is running and when it ends:
[yourUsername@login1 ~]$ squeue -u yourUsername
Use ls to locate the output file. The -t flag sorts in
reverse-chronological order: newest first. What was the output?
Read the Job Output
The cluster output should be written to a file in the folder you launched the job from. For example,
[yourUsername@login1 ~]$ ls -tslurm-347087.out serial-job.sh amdahl README.md LICENSE.txt[yourUsername@login1 ~]$ cat slurm-347087.outDoing 30.000 seconds of 'work' on 1 processor, which should take 30.000 seconds with 0.850 parallel proportion of the workload. Hello, World! I am process 0 of 1 on node1. I will do all the serial 'work' for 4.500 seconds. Hello, World! I am process 0 of 1 on node1. I will do parallel 'work' for 25.500 seconds. Total execution time (according to rank 0): 30.033 seconds
As we saw before, two of the amdahl program flags set the amount of work and
the proportion of that work that is parallel in nature. Based on the output, we
can see that the code uses a default of 30 seconds of work that is 85%
parallel. The program ran for just over 30 seconds in total, and if we run the
numbers, it is true that 15% of it was marked ‘serial’ and 85% was ‘parallel’.
Since we only gave the job one CPU, this job wasn’t really parallel: the same processor performed the ‘serial’ work for 4.5 seconds, then the ‘parallel’ part for 25.5 seconds, and no time was saved. The cluster can do better, if we ask.
Running the Parallel Job
The amdahl program uses the Message Passing Interface (MPI) for parallelism
– this is a common tool on HPC systems.
What is MPI?
The Message Passing Interface is a set of tools which allow multiple tasks running simultaneously to communicate with each other. Typically, a single executable is run multiple times, possibly on different machines, and the MPI tools are used to inform each instance of the executable about its sibling processes, and which instance it is. MPI also provides tools to allow communication between instances to coordinate work, exchange information about elements of the task, or to transfer data. An MPI instance typically has its own copy of all the local variables.
While MPI-aware executables can generally be run as stand-alone programs, in
order for them to run in parallel they must use an MPI run-time environment,
which is a specific implementation of the MPI standard.
To activate the MPI environment, the program should be started via a command
such as mpiexec (or mpirun, or srun, etc. depending on the MPI run-time
you need to use), which will ensure that the appropriate run-time support for
parallelism is included.
MPI Runtime Arguments
On their own, commands such as
mpiexeccan take many arguments specifying how many machines will participate in the execution, and you might need these if you would like to run an MPI program on your own (for example, on your laptop). In the context of a queuing system, however, it is frequently the case that MPI run-time will obtain the necessary parameters from the queuing system, by examining the environment variables set when the job is launched.
Let’s modify the job script to request more cores and use the MPI run-time.
[yourUsername@login1 ~]$ cp serial-job.sh parallel-job.sh
[yourUsername@login1 ~]$ nano parallel-job.sh
[yourUsername@login1 ~]$ cat parallel-job.sh
#!/bin/bash
#SBATCH -J parallel-job
#SBATCH -p cpubase_bycore_b1
#SBATCH -N 1
#SBATCH -n 4
# Load the computing environment we need
# (mpi4py and numpy are in SciPy-bundle)
module load Python
module load SciPy-bundle
# Execute the task
mpiexec amdahl
Then submit your job. Note that the submission command has not really changed from how we submitted the serial job: all the parallel settings are in the batch file rather than the command line.
[yourUsername@login1 ~]$ sbatch parallel-job.sh
As before, use the status commands to check when your job runs.
[yourUsername@login1 ~]$ ls -t
slurm-347178.out parallel-job.sh slurm-347087.out serial-job.sh amdahl README.md LICENSE.txt
[yourUsername@login1 ~]$ cat slurm-347178.out
Doing 30.000 seconds of 'work' on 4 processors,
which should take 10.875 seconds with 0.850 parallel proportion of the workload.
Hello, World! I am process 0 of 4 on node1. I will do all the serial 'work' for 4.500 seconds.
Hello, World! I am process 2 of 4 on node1. I will do parallel 'work' for 6.375 seconds.
Hello, World! I am process 1 of 4 on node1. I will do parallel 'work' for 6.375 seconds.
Hello, World! I am process 3 of 4 on node1. I will do parallel 'work' for 6.375 seconds.
Hello, World! I am process 0 of 4 on node1. I will do parallel 'work' for 6.375 seconds.
Total execution time (according to rank 0): 10.888 seconds
Is it 4× faster?
The parallel job received 4× more processors than the serial job: does that mean it finished in ¼ the time?
Solution
The parallel job did take less time: 11 seconds is better than 30! But it is only a 2.7× improvement, not 4×.
Look at the job output:
- While “process 0” did serial work, processes 1 through 3 did their parallel work.
- While process 0 caught up on its parallel work, the rest did nothing at all.
Process 0 always has to finish its serial task before it can start on the parallel work. This sets a lower limit on the amount of time this job will take, no matter how many cores you throw at it.
This is the basic principle behind Amdahl’s Law, which is one way of predicting improvements in execution time for a fixed workload that can be subdivided and run in parallel to some extent.
How Much Does Parallel Execution Improve Performance?
In theory, dividing up a perfectly parallel calculation among n MPI processes should produce a decrease in total run time by a factor of n. As we have just seen, real programs need some time for the MPI processes to communicate and coordinate, and some types of calculations can’t be subdivided: they only run effectively on a single CPU.
Additionally, if the MPI processes operate on different physical CPUs in the computer, or across multiple compute nodes, even more time is required for communication than it takes when all processes operate on a single CPU.
In practice, it’s common to evaluate the parallelism of an MPI program by
- running the program across a range of CPU counts,
- recording the execution time on each run,
- comparing each execution time to the time when using a single CPU.
Since “more is better” – improvement is easier to interpret from increases in some quantity than decreases – comparisons are made using the speedup factor S, which is calculated as the single-CPU execution time divided by the multi-CPU execution time. For a perfectly parallel program, a plot of the speedup S versus the number of CPUs n would give a straight line, S = n.
Let’s run one more job, so we can see how close to a straight line our amdahl
code gets.
[yourUsername@login1 ~]$ nano parallel-job.sh
[yourUsername@login1 ~]$ cat parallel-job.sh
#!/bin/bash
#SBATCH -J parallel-job
#SBATCH -p cpubase_bycore_b1
#SBATCH -N 1
#SBATCH -n 8
# Load the computing environment we need
# (mpi4py and numpy are in SciPy-bundle)
module load Python
module load SciPy-bundle
# Execute the task
mpiexec amdahl
Then submit your job. Note that the submission command has not really changed from how we submitted the serial job: all the parallel settings are in the batch file rather than the command line.
[yourUsername@login1 ~]$ sbatch parallel-job.sh
As before, use the status commands to check when your job runs.
[yourUsername@login1 ~]$ ls -t
slurm-347271.out parallel-job.sh slurm-347178.out slurm-347087.out serial-job.sh amdahl README.md LICENSE.txt
[yourUsername@login1 ~]$ cat slurm-347178.out
which should take 7.688 seconds with 0.850 parallel proportion of the workload.
Hello, World! I am process 4 of 8 on node1. I will do parallel 'work' for 3.188 seconds.
Hello, World! I am process 0 of 8 on node1. I will do all the serial 'work' for 4.500 seconds.
Hello, World! I am process 2 of 8 on node1. I will do parallel 'work' for 3.188 seconds.
Hello, World! I am process 1 of 8 on node1. I will do parallel 'work' for 3.188 seconds.
Hello, World! I am process 3 of 8 on node1. I will do parallel 'work' for 3.188 seconds.
Hello, World! I am process 5 of 8 on node1. I will do parallel 'work' for 3.188 seconds.
Hello, World! I am process 6 of 8 on node1. I will do parallel 'work' for 3.188 seconds.
Hello, World! I am process 7 of 8 on node1. I will do parallel 'work' for 3.188 seconds.
Hello, World! I am process 0 of 8 on node1. I will do parallel 'work' for 3.188 seconds.
Total execution time (according to rank 0): 7.697 seconds
Non-Linear Output
When we ran the job with 4 parallel workers, the serial job wrote its output first, then the parallel processes wrote their output, with process 0 coming in first and last.
With 8 workers, this is not the case: since the parallel workers take less time than the serial work, it is hard to say which process will write its output first, except that it will not be process 0!
Now, let’s summarize the amount of time it took each job to run:
| Number of CPUs | Runtime (sec) |
|---|---|
| 1 | 30.033 |
| 4 | 10.888 |
| 8 | 7.697 |
Then, use the first row to compute speedups S, using Python as a command-line calculator:
[yourUsername@login1 ~]$ for n in 30.033 10.888 7.697; do python3 -c "print(30.033 / $n)"; done
| Number of CPUs | Speedup | Ideal |
|---|---|---|
| 1 | 1.0 | 1 |
| 4 | 2.75 | 4 |
| 8 | 3.90 | 8 |
The job output files have been telling us that this program is performing 85% of its work in parallel, leaving 15% to run in serial. This seems reasonably high, but our quick study of speedup shows that in order to get a 4× speedup, we have to use 8 or 9 processors in parallel. In real programs, the speedup factor is influenced by
- CPU design
- communication network between compute nodes
- MPI library implementations
- details of the MPI program itself
Using Amdahl’s Law, you can prove that with this program, it is impossible to reach 8× speedup, no matter how many processors you have on hand. Details of that analysis, with results to back it up, are left for the next class in the HPC Carpentry workshop, HPC Workflows.
In an HPC environment, we try to reduce the execution time for all types of jobs, and MPI is an extremely common way to combine dozens, hundreds, or thousands of CPUs into solving a single problem. To learn more about parallelization, see the parallel novice lesson lesson.
Key Points
Parallel programming allows applications to take advantage of parallel hardware.
The queuing system facilitates executing parallel tasks.
Performance improvements from parallel execution do not scale linearly.
Using resources effectively
Overview
Teaching: 10 min
Exercises: 20 minQuestions
How can I review past jobs?
How can I use this knowledge to create a more accurate submission script?
Objectives
Look up job statistics.
Make more accurate resource requests in job scripts based on data describing past performance.
We’ve touched on all the skills you need to interact with an HPC cluster: logging in over SSH, loading software modules, submitting parallel jobs, and finding the output. Let’s learn about estimating resource usage and why it might matter.
Estimating Required Resources Using the Scheduler
Although we covered requesting resources from the scheduler earlier with the π code, how do we know what type of resources the software will need in the first place, and its demand for each? In general, unless the software documentation or user testimonials provide some idea, we won’t know how much memory or compute time a program will need.
Read the Documentation
Most HPC facilities maintain documentation as a wiki, a website, or a document sent along when you register for an account. Take a look at these resources, and search for the software you plan to use: somebody might have written up guidance for getting the most out of it.
A convenient way of figuring out the resources required for a job to run
successfully is to submit a test job, and then ask the scheduler about its
impact using sacct -u yourUsername. You can use this knowledge to set up the
next job with a closer estimate of its load on the system. A good general rule
is to ask the scheduler for 20% to 30% more time and memory than you expect the
job to need. This ensures that minor fluctuations in run time or memory use
will not result in your job being cancelled by the scheduler. Keep in mind that
if you ask for too much, your job may not run even though enough resources are
available, because the scheduler will be waiting for other people’s jobs to
finish and free up the resources needed to match what you asked for.
Stats
Since we already submitted amdahl to run on the cluster, we can query the
scheduler to see how long our job took and what resources were used. We will
use sacct -u yourUsername to get statistics about parallel-job.sh.
[yourUsername@login1 ~]$ sacct -u yourUsername
JobID JobName Partition Account AllocCPUS State ExitCode
------------ ---------- ---------- ---------- ---------- ---------- --------
7 file.sh cpubase_b+ def-spons+ 1 COMPLETED 0:0
7.batch batch def-spons+ 1 COMPLETED 0:0
7.extern extern def-spons+ 1 COMPLETED 0:0
8 file.sh cpubase_b+ def-spons+ 1 COMPLETED 0:0
8.batch batch def-spons+ 1 COMPLETED 0:0
8.extern extern def-spons+ 1 COMPLETED 0:0
9 example-j+ cpubase_b+ def-spons+ 1 COMPLETED 0:0
9.batch batch def-spons+ 1 COMPLETED 0:0
9.extern extern def-spons+ 1 COMPLETED 0:0
This shows all the jobs we ran today (note that there are multiple entries per job). To get info about a specific job (for example, 347087), we change command slightly.
[yourUsername@login1 ~]$ sacct -u yourUsername -l -j 347087
It will show a lot of info; in fact, every single piece of info collected on
your job by the scheduler will show up here. It may be useful to redirect this
information to less to make it easier to view (use the left and right arrow
keys to scroll through fields).
[yourUsername@login1 ~]$ sacct -u yourUsername -l -j 347087 | less -S
Discussion
This view can help compare the amount of time requested and actually used, duration of residence in the queue before launching, and memory footprint on the compute node(s).
How accurate were our estimates?
Improving Resource Requests
From the job history, we see that amdahl jobs finished executing in
at most a few minutes, once dispatched. The time estimate we provided
in the job script was far too long! This makes it harder for the
queuing system to accurately estimate when resources will become free
for other jobs. Practically, this means that the queuing system waits
to dispatch our amdahl job until the full requested time slot opens,
instead of “sneaking it in” a much shorter window where the job could
actually finish. Specifying the expected runtime in the submission
script more accurately will help alleviate cluster congestion and may
get your job dispatched earlier.
Narrow the Time Estimate
Edit
parallel_job.shto set a better time estimate. How close can you get?Hint: use
-t.Solution
The following line tells Slurm that our job should finish within 2 minutes:
#SBATCH -t 00:02:00
Key Points
Accurate job scripts help the queuing system efficiently allocate shared resources.
Using shared resources responsibly
Overview
Teaching: 15 min
Exercises: 5 minQuestions
How can I be a responsible user?
How can I protect my data?
How can I best get large amounts of data off an HPC system?
Objectives
Describe how the actions of a single user can affect the experience of others on a shared system.
Discuss the behaviour of a considerate shared system citizen.
Explain the importance of backing up critical data.
Describe the challenges with transferring large amounts of data off HPC systems.
Convert many files to a single archive file using tar.
One of the major differences between using remote HPC resources and your own system (e.g. your laptop) is that remote resources are shared. How many users the resource is shared between at any one time varies from system to system, but it is unlikely you will ever be the only user logged into or using such a system.
The widespread usage of scheduling systems where users submit jobs on HPC resources is a natural outcome of the shared nature of these resources. There are other things you, as an upstanding member of the community, need to consider.
Be Kind to the Login Nodes
The login node is often busy managing all of the logged in users, creating and editing files and compiling software. If the machine runs out of memory or processing capacity, it will become very slow and unusable for everyone. While the machine is meant to be used, be sure to do so responsibly – in ways that will not adversely impact other users’ experience.
Login nodes are always the right place to launch jobs. Cluster policies vary, but they may also be used for proving out workflows, and in some cases, may host advanced cluster-specific debugging or development tools. The cluster may have modules that need to be loaded, possibly in a certain order, and paths or library versions that differ from your laptop, and doing an interactive test run on the head node is a quick and reliable way to discover and fix these issues.
Login Nodes Are a Shared Resource
Remember, the login node is shared with all other users and your actions could cause issues for other people. Think carefully about the potential implications of issuing commands that may use large amounts of resource.
Unsure? Ask your friendly systems administrator (“sysadmin”) if the thing you’re contemplating is suitable for the login node, or if there’s another mechanism to get it done safely.
You can always use the commands top and ps ux to list the processes that
are running on the login node along with the amount of CPU and memory they are
using. If this check reveals that the login node is somewhat idle, you can
safely use it for your non-routine processing task. If something goes wrong
– the process takes too long, or doesn’t respond – you can use the
kill command along with the PID to terminate the process.
Login Node Etiquette
Which of these commands would be a routine task to run on the login node?
python physics_sim.pymakecreate_directories.shmolecular_dynamics_2tar -xzf R-3.3.0.tar.gzSolution
Building software, creating directories, and unpacking software are common and acceptable > tasks for the login node: options #2 (
make), #3 (mkdir), and #5 (tar) are probably OK. Note that script names do not always reflect their contents: before launching #3, pleaseless create_directories.shand make sure it’s not a Trojan horse.Running resource-intensive applications is frowned upon. Unless you are sure it will not affect other users, do not run jobs like #1 (
python) or #4 (custom MD code). If you’re unsure, ask your friendly sysadmin for advice.
If you experience performance issues with a login node you should report it to the system staff (usually via the helpdesk) for them to investigate.
Test Before Scaling
Remember that you are generally charged for usage on shared systems. A simple mistake in a job script can end up costing a large amount of resource budget. Imagine a job script with a mistake that makes it sit doing nothing for 24 hours on 1000 cores or one where you have requested 2000 cores by mistake and only use 100 of them! This problem can be compounded when people write scripts that automate job submission (for example, when running the same calculation or analysis over lots of different parameters or files). When this happens it hurts both you (as you waste lots of charged resource) and other users (who are blocked from accessing the idle compute nodes). On very busy resources you may wait many days in a queue for your job to fail within 10 seconds of starting due to a trivial typo in the job script. This is extremely frustrating!
Most systems provide dedicated resources for testing that have short wait times to help you avoid this issue.
Test Job Submission Scripts That Use Large Amounts of Resources
Before submitting a large run of jobs, submit one as a test first to make sure everything works as expected.
Before submitting a very large or very long job submit a short truncated test to ensure that the job starts as expected.
Have a Backup Plan
Although many HPC systems keep backups, it does not always cover all the file systems available and may only be for disaster recovery purposes (i.e. for restoring the whole file system if lost rather than an individual file or directory you have deleted by mistake). Protecting critical data from corruption or deletion is primarily your responsibility: keep your own backup copies.
Version control systems (such as Git) often have free, cloud-based offerings (e.g., GitHub and GitLab) that are generally used for storing source code. Even if you are not writing your own programs, these can be very useful for storing job scripts, analysis scripts and small input files.
If you are building software, you may have a large amount of source code that you compile to build your executable. Since this data can generally be recovered by re-downloading the code, or re-running the checkout operation from the source code repository, this data is also less critical to protect.
For larger amounts of data, especially important results from your runs,
which may be irreplaceable, you should make sure you have a robust system in
place for taking copies of data off the HPC system wherever possible
to backed-up storage. Tools such as rsync can be very useful for this.
Your access to the shared HPC system will generally be time-limited so you should ensure you have a plan for transferring your data off the system before your access finishes. The time required to transfer large amounts of data should not be underestimated and you should ensure you have planned for this early enough (ideally, before you even start using the system for your research).
In all these cases, the helpdesk of the system you are using should be able to provide useful guidance on your options for data transfer for the volumes of data you will be using.
Your Data Is Your Responsibility
Make sure you understand what the backup policy is on the file systems on the system you are using and what implications this has for your work if you lose your data on the system. Plan your backups of critical data and how you will transfer data off the system throughout the project.
Transferring Data
As mentioned above, many users run into the challenge of transferring large amounts of data off HPC systems at some point (this is more often in transferring data off than onto systems but the advice below applies in either case). Data transfer speed may be limited by many different factors so the best data transfer mechanism to use depends on the type of data being transferred and where the data is going.
The components between your data’s source and destination have varying levels of performance, and in particular, may have different capabilities with respect to bandwidth and latency.
Bandwidth is generally the raw amount of data per unit time a device is capable of transmitting or receiving. It’s a common and generally well-understood metric.
Latency is a bit more subtle. For data transfers, it may be thought of as the amount of time it takes to get data out of storage and into a transmittable form. Latency issues are the reason it’s advisable to execute data transfers by moving a small number of large files, rather than the converse.
Some of the key components and their associated issues are:
- Disk speed: File systems on HPC systems are often highly parallel, consisting of a very large number of high performance disk drives. This allows them to support a very high data bandwidth. Unless the remote system has a similar parallel file system you may find your transfer speed limited by disk performance at that end.
- Meta-data performance: Meta-data operations such as opening and closing files or listing the owner or size of a file are much less parallel than read/write operations. If your data consists of a very large number of small files you may find your transfer speed is limited by meta-data operations. Meta-data operations performed by other users of the system can also interact strongly with those you perform so reducing the number of such operations you use (by combining multiple files into a single file) may reduce variability in your transfer rates and increase transfer speeds.
- Network speed: Data transfer performance can be limited by network speed. More importantly it is limited by the slowest section of the network between source and destination. If you are transferring to your laptop/workstation, this is likely to be its connection (either via LAN or WiFi).
- Firewall speed: Most modern networks are protected by some form of firewall that filters out malicious traffic. This filtering has some overhead and can result in a reduction in data transfer performance. The needs of a general purpose network that hosts email/web-servers and desktop machines are quite different from a research network that needs to support high volume data transfers. If you are trying to transfer data to or from a host on a general purpose network you may find the firewall for that network will limit the transfer rate you can achieve.
As mentioned above, if you have related data that consists of a large number of
small files it is strongly recommended to pack the files into a larger
archive file for long term storage and transfer. A single large file makes
more efficient use of the file system and is easier to move, copy and transfer
because significantly fewer metadata operations are required. Archive files can
be created using tools like tar and zip. We have already met tar when we
talked about data transfer earlier.
Consider the Best Way to Transfer Data
If you are transferring large amounts of data you will need to think about what may affect your transfer performance. It is always useful to run some tests that you can use to extrapolate how long it will take to transfer your data.
Say you have a “data” folder containing 10,000 or so files, a healthy mix of small and large ASCII and binary data. Which of the following would be the best way to transfer them to HPC Carpentry’s Cloud Cluster?
[you@laptop:~]$ scp -r data yourUsername@cluster.hpc-carpentry.org:~/[you@laptop:~]$ rsync -ra data yourUsername@cluster.hpc-carpentry.org:~/[you@laptop:~]$ rsync -raz data yourUsername@cluster.hpc-carpentry.org:~/[you@laptop:~]$ tar -cvf data.tar data [you@laptop:~]$ rsync -raz data.tar yourUsername@cluster.hpc-carpentry.org:~/[you@laptop:~]$ tar -cvzf data.tar.gz data [you@laptop:~]$ rsync -ra data.tar.gz yourUsername@cluster.hpc-carpentry.org:~/Solution
scpwill recursively copy the directory. This works, but without compression.rsync -raworks likescp -r, but preserves file information like creation times. This is marginally better.rsync -razadds compression, which will save some bandwidth. If you have a strong CPU at both ends of the line, and you’re on a slow network, this is a good choice.- This command first uses
tarto merge everything into a single file, thenrsync -zto transfer it with compression. With this large number of files, metadata overhead can hamper your transfer, so this is a good idea.- This command uses
tar -zto compress the archive, thenrsyncto transfer it. This may perform similarly to #4, but in most cases (for large datasets), it’s the best combination of high throughput and low latency (making the most of your time and network connection).
Key Points
Be careful how you use the login node.
Your data on the system is your responsibility.
Plan and test large data transfers.
It is often best to convert many files to a single archive file before transferring.